Figures & data
Figure 1. (A) Bar diagram of total leaf proteins with standard errors from the plants grown in saline and non-saline habitat. (B) SDS-PAGE documentation of five taxa grown in saline and non-saline habitat; in the pairs of lanes, the left lane represents saline and the right one forms non-saline habitat plants. M – Protein marker, 1 – B. gymnorrhiza, 2 – E. agallocha, 3 – H. fomes, 4 – P. paludosa and 5 – X. granatum.
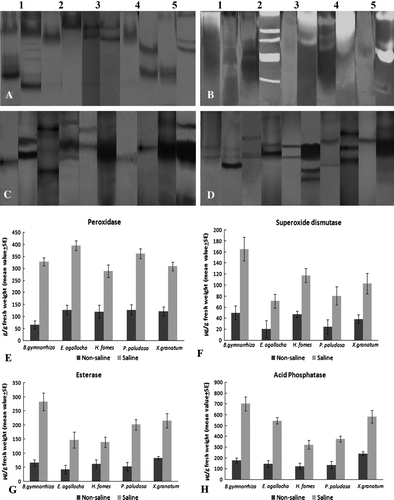
Figure 2. (A–D) Documented native gel electrophoresis. (A) Peroxidase; (B) Superoxide dismutase; (C) Esterase; (D) Acid phosphatase. In all cases, left lane from non-saline and right one from saline plants. 1 – B. gymnorrhiza, 2 – E. agallocha, 3 – H. fomes, 4 – P. paludosa and 5 – X. granatum. (E–H) Bar diagrams of quantitative estimation of enzymes with standard errors bars.