Figures & data
Figure 2. XIC of modified and unmodified tryptic peptide XXXXXXXXXWGQGTLVTVSSASTK. (Note the ∼100-fold difference in intensity scale between upper and lower traces.) Peaks 1a and 1b correspond to tryptophan oxidation species that are isobaric with the modified peptide of peak 3.
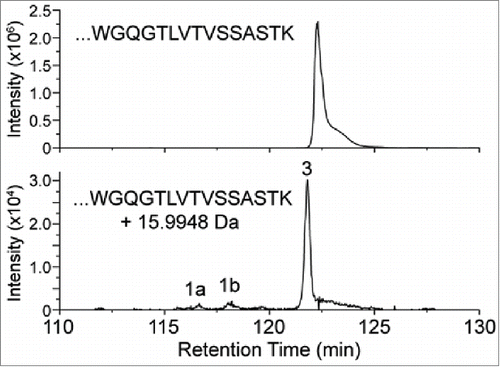
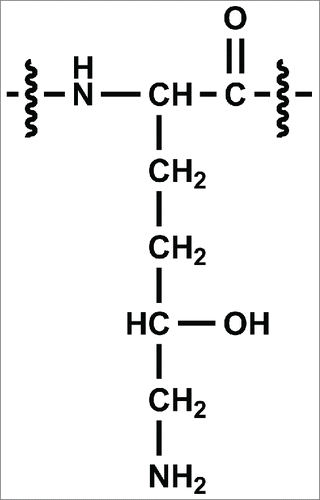
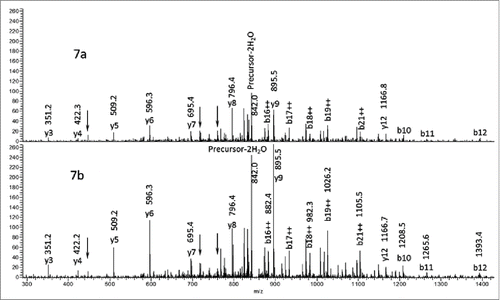
Figure 3. CID data from the modified (3a) m/z 854.1 [M+3H]3+ peptide and unmodified (3b) peptide m/z 848.8 [M+3H]3+. The oxygen addition is localized to one of the amino acids in bold.
![Figure 3. CID data from the modified (3a) m/z 854.1 [M+3H]3+ peptide and unmodified (3b) peptide m/z 848.8 [M+3H]3+. The oxygen addition is localized to one of the amino acids in bold.](/cms/asset/e99f58a7-d9cc-4053-98b2-a00db6718a0e/kmab_a_1122148_f0003_b.gif)
Figure 4. UPLC-MS extracted ion chromatogram of m/z 854.0789 [M+3H]3+ of tryptic digest from mAb1 spiked with a synthetic peptide containing alanine to serine sequence variant (identified as peak 2). The other identified peaks (1a , 1b and 4) correspond to peptides containing isomers of oxidized tryptophan at position 10 in the peptide, and the potential lysine to hydroxylysine modification at the C-terminus (peak 3).
![Figure 4. UPLC-MS extracted ion chromatogram of m/z 854.0789 [M+3H]3+ of tryptic digest from mAb1 spiked with a synthetic peptide containing alanine to serine sequence variant (identified as peak 2). The other identified peaks (1a , 1b and 4) correspond to peptides containing isomers of oxidized tryptophan at position 10 in the peptide, and the potential lysine to hydroxylysine modification at the C-terminus (peak 3).](/cms/asset/0cad147d-cfd5-4afe-b932-25dd36209431/kmab_a_1122148_f0004_b.gif)
Figure 5. MS3 of tryptic Hyl-containing (5a) and unmodified (5b) peptides. Precursor3+ (y3)1+ Fragments.
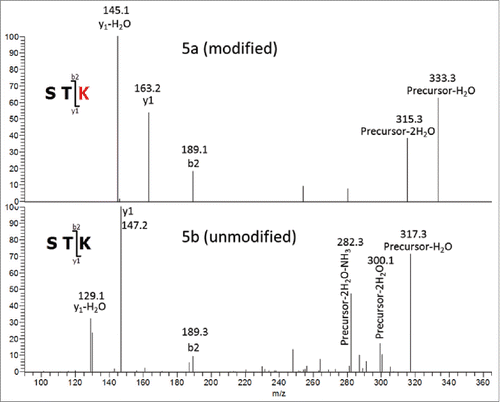
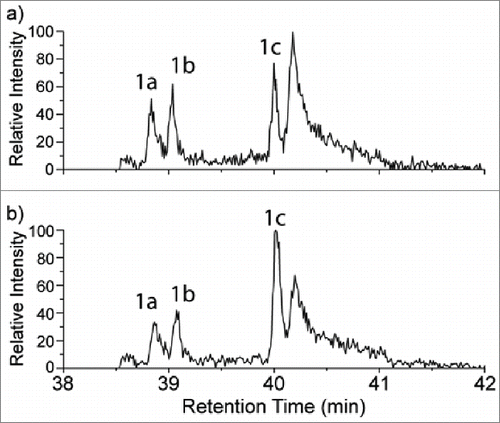
Figure 6. A portion of the extracted ion chromatograms (XICs) at m/z 854.0789 for the mAb1 tryptic digests both without (6a) and with (6b) addition of the synthetic Hyl-containing peptide. Peak 1c in each chromatogram corresponds to the retention times of both the native +16 Da peptide (6a) and the combined native and spiked Hyl-containing peptide (6b). Peaks 1a and 1b were identified as the oxidized tryptophan form of the peptide, which is isobaric with the Hyl-containing peptide.