Figures & data
Figure 1. Deconvoluted mass spectrum of denatured and reduced Fc-G4P-protein. The Fc is linked to the protein via a G4P linker. Typical Fc glycosylation (G0F, G1F, G2F, etc.) was observed. Modification #1 represents about 45% of the MS signal based on peak height.
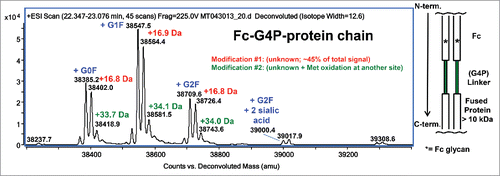
Figure 2. A high level +15.99 Da mass increase was observed on the G4P linker peptide by LC-MS/MS. Based on peptide signal intensity, there is a ratio of about 60:40 of unmodified peptide: modified peptide. Top panel: XIC (Extracted Ion Chromatogram) of unmodified and modified peptides. Bottom panel: Centroid mass spectrum averaged from T21.5–24.5 min.
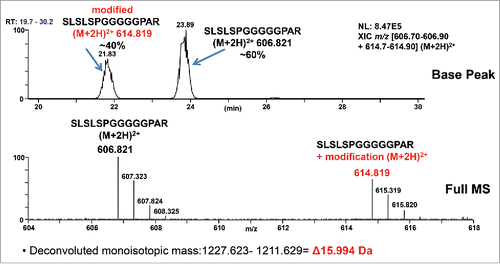
Figure 3. Low-resolution CID mass spectra of unmodified and +15.99 Da modified ([M+2H]2+ = 614.82) linker peptide SLSLSPGGGGGPAR. The modification can be localized to the GGGPAR portion of the sequence by the difference in y6 ions. Different b13 ions eliminate the C-terminal R.
![Figure 3. Low-resolution CID mass spectra of unmodified and +15.99 Da modified ([M+2H]2+ = 614.82) linker peptide SLSLSPGGGGGPAR. The modification can be localized to the GGGPAR portion of the sequence by the difference in y6 ions. Different b13 ions eliminate the C-terminal R.](/cms/asset/9742d882-5a4a-4945-872c-8411846e533f/kmab_a_1325556_f0003_c.gif)
Figure 4. High-resolution HCD mass spectrum of +15.99 Da modified ([M+2H]2+ = 614.82) linker peptide SLSLSPGGGGGPAR. Top panel: m/z [100–1250]. Bottom panel: m/z [340–460]. The detected accurate masses of the y3 and y4 fragment ions rule out Pro221→ Leu/Ile221 mutation. The HCD fragmentation cannot discriminate between Pro221→Hyp221 or Ala222→Ser222 as no y2 fragment ion was observed.
![Figure 4. High-resolution HCD mass spectrum of +15.99 Da modified ([M+2H]2+ = 614.82) linker peptide SLSLSPGGGGGPAR. Top panel: m/z [100–1250]. Bottom panel: m/z [340–460]. The detected accurate masses of the y3 and y4 fragment ions rule out Pro221→ Leu/Ile221 mutation. The HCD fragmentation cannot discriminate between Pro221→Hyp221 or Ala222→Ser222 as no y2 fragment ion was observed.](/cms/asset/eea9062e-ebe8-4f1c-ac94-c37e3b969121/kmab_a_1325556_f0004_c.gif)
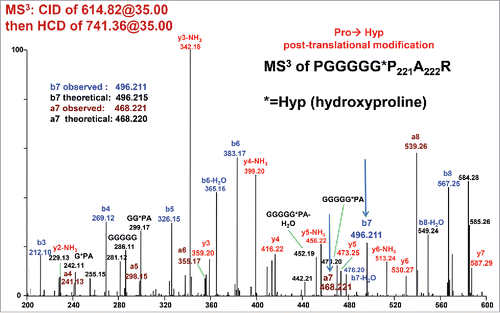
Figure 5. CID/HCD MS3 spectrum of m/z 741.36, i.e. y9 ion resulting from CID fragmentation of the +15.99 Da modified peptide (see Bottom Panel of ). HCD fragmentation of the 741.36 ion yields unique a7 and b7 fragment ions that confirm a Pro221→Hyp 221 post-translational modification.

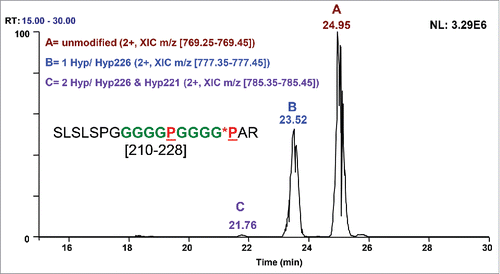
![Figure 7. High-resolution CID mass spectrum of the doubly charged ion of (G4P)2 linker peptide SLSLSPGGGGGPGGGGPAR [210–228] with one Hyp. Hyp PTM is present in y3 and higher y ions, confirming Hyp226.](/cms/asset/ebf76b52-1245-4ef7-808a-80345606525f/kmab_a_1325556_f0007_c.gif)