Figures & data
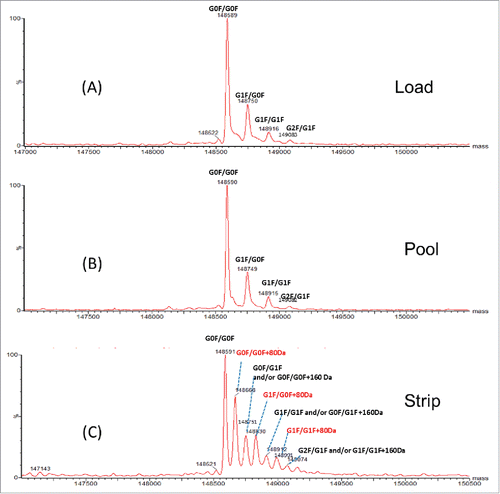
Figure 1. Overlay of SCX-HPLC UV profile of AEX load (blue trace), strip (red trace) and pool fraction (green trace).
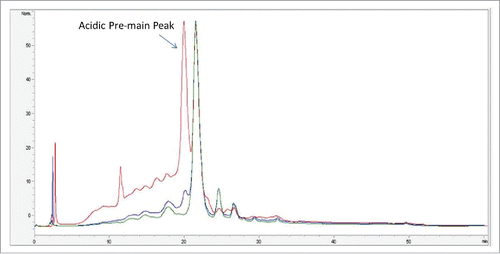
Figure 5. (A) CID fragmentation spectrum of light chain AA25–43+80 Da in 400–1800 m/z (B) Zoom in m/z 300–1100 (C) Zoomed in m/z 1200–2000.
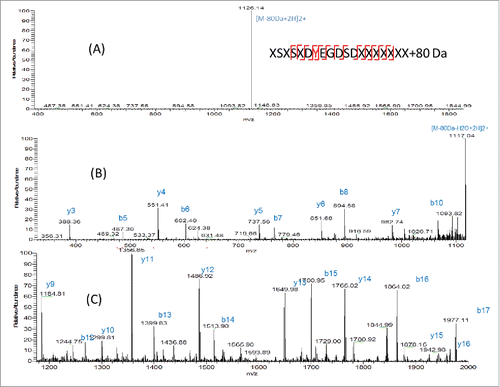
Figure 6. ETD fragmentation of light chain peptide AA25–43+80 Da. The 80 Da attached fragment ions were highlighted in red and labeled with *.
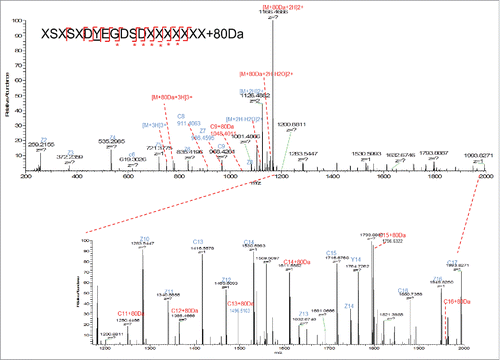
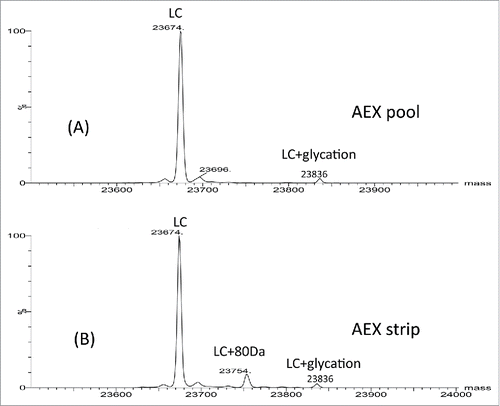
Figure 7. (A) Deconvoluted intact mass spectra of AEX strip fraction with and without alkaline phosphatase treatment. (B) Deconvoluted intact mass spectra of chicken ovalbumin with and without alkaline phosphatase treatment.
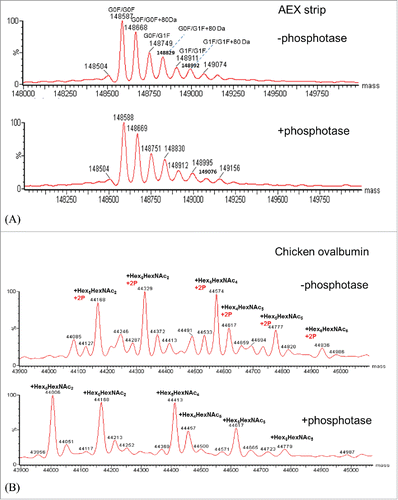
Figure 8. (a) Normalized concentrations of mAb AEX pool and strip were subjected to reduced SDS-PAGE, probed for the human heavy (HC) and light chains (LC) by western hybridization (upper panel), then stripped and re-probed for antisulfotyrosine (lower panel). See the indications for HC and LC at the far right. (b) Normalized concentrations of different CHO-derived mAbs in addition to AEX strip and pool are subjected to reduced SDS PAGE, probed for the human HC and LC by western hybridization, then stripped and re-probed for anti sulfotyrosine. For both (a) and (b) MagicMark XP was used as a protein molecular weight standard, and equal amounts of HEK293 and EGF-treated A431 cell extracts are analyzed as controls.
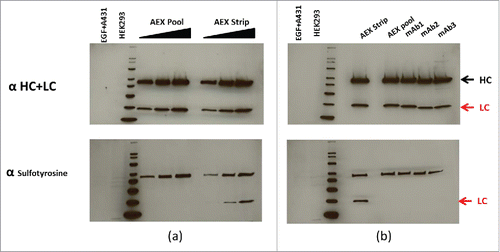
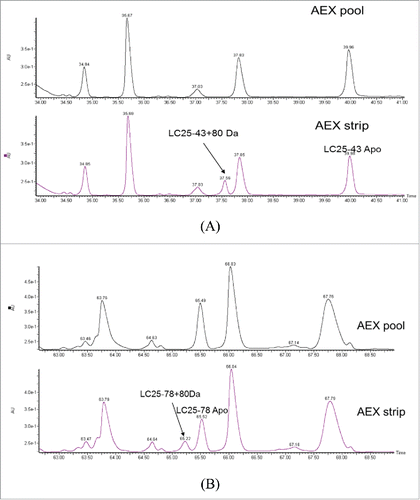
Figure 9. SIC of (A) LC25–43C80 Da from AEX Strip Fraction, (B) Synthetic peptide XSXSXDYEGDSDXXXXXXXCPhosphorylation and (C) Synthetic peptide XSXSXDYEGDSDXXXXXXXCSulfation; ETD MS2 spectra of (D) LC25–43C80 Da from AEX strip fraction and (E) Synthetic peptide XSXSXDYEGDSDXXXXXXXCSulfation.
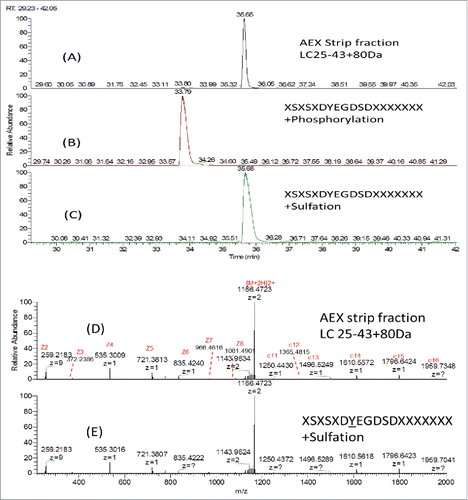
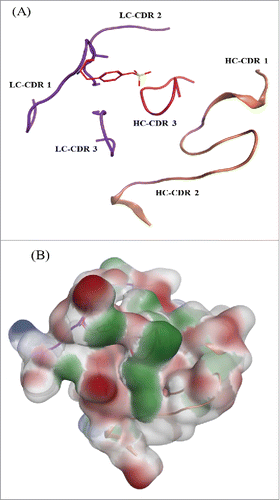
Figure 10. mAb tyrosine (Y31) site showing the CDR loops for both the heavy and light chain. (A) Sulfated tyrosine (Try31) is depicted in red sticks and in the context of the CDR regions in ribbon diagram. (B) Surface map of the CDR region shown in the same orientation as the ribbon diagram. Hydrophobic regions in green, negatively charges regions in red, and positively charged region in blue.