Figures & data
Table 1. Cross-industry studies demonstrating the application LC-MS/MS methods in support of process development.
Table 2. Recent studies and trend of LC-MS based analysis in HCP characterization.
Table 3. Comparison of different LC and MS methods with their advantages and limitations for HCP analysis.
Figure 1. General workflow for HCP characterization by LC-MS/MS analysis.
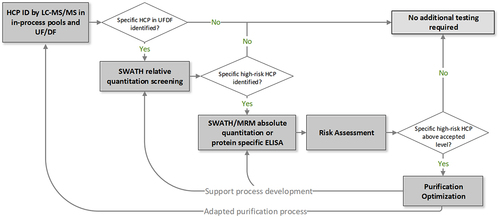
Table 4. Our commonly used HCP identification methods by LC-MS/MS analysis.
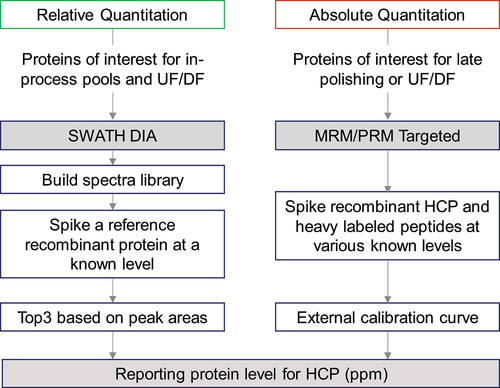
Figure 2. Workflow for HCP quantitation by SWATH and targeted (MRM/PRM) MS analysis.