Figures & data
Figure 1. Binding epitopes of antigen-antibody pairs as determined by different technologies.
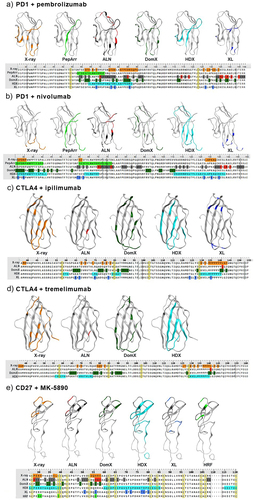
Figure 2. Antibody binding to mouse/human domain exchange mutants.
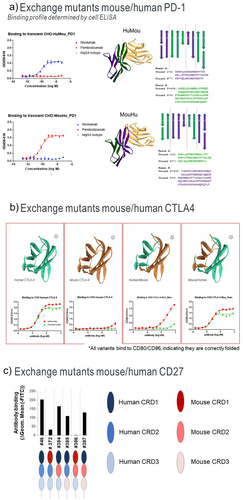
Figure 3. Binding of anti-CTLA-4 antibodies to alanine mutants.
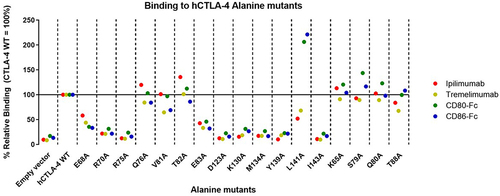
Table 1. Pro and cons of each technique.
