Figures & data
Table 1. Characteristics of study participants.
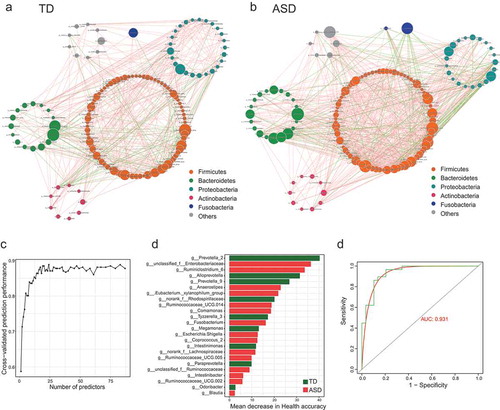
Figure 1. The shift of gut microbiota in typically developing (TD) and ASD children according to the 16S rRNA data.
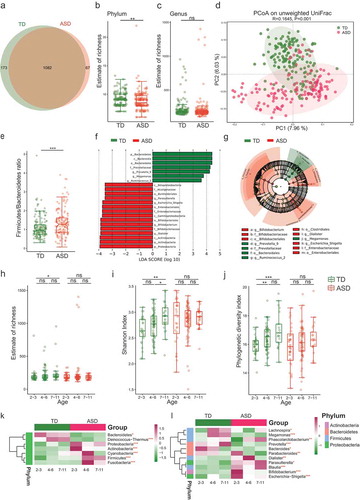
Figure 3. The gut microbiota divergence in ASD with constipation (C-ASD) and TD children based on the metagenomic sequencing data.
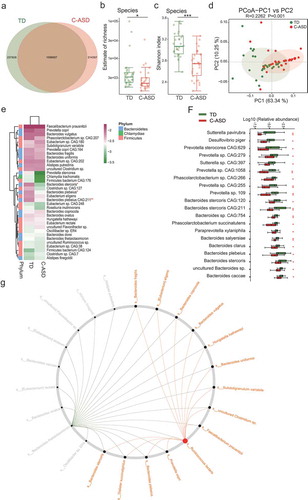
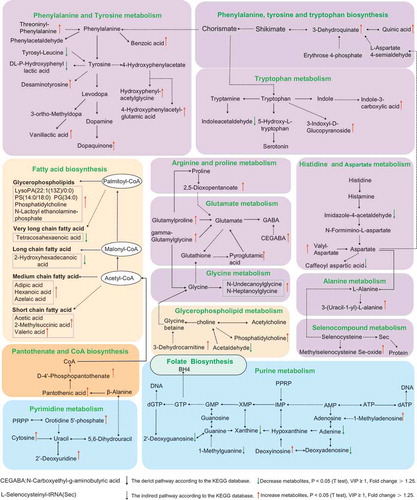
Figure 4. Microbial gene functions annotation on KEGG in ASD with constipation (C-ASD) and TD children.
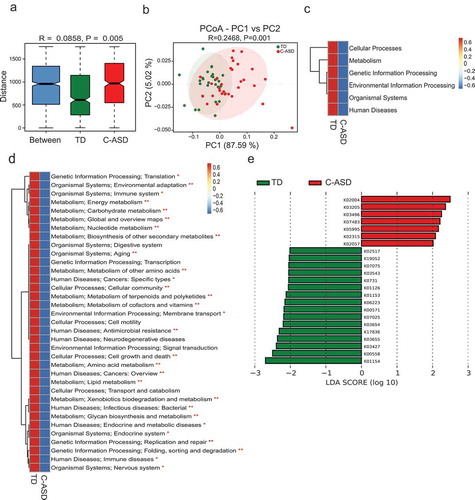
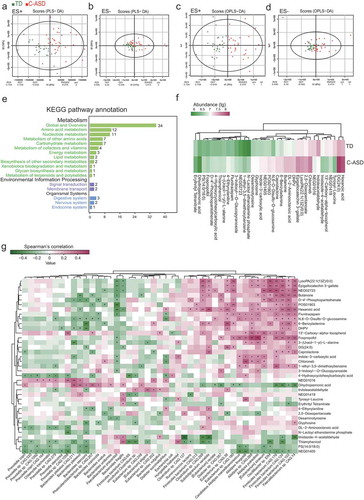
Figure 5. Aberrant metabolic patterns in ASD with constipation (C-ASD) and typically developing (TD) children.