Figures & data
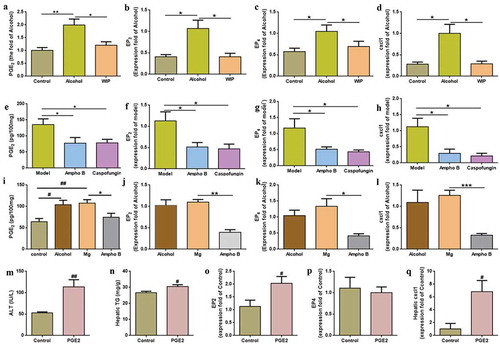
Figure 1. Oral treatment with WIP alleviates chronic ethanol feeding-induced hepatic injury and steatosis
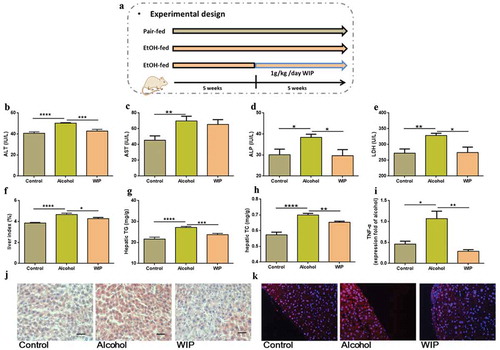
Figure 2. WIP treatment ameliorates the ethanol-induced gut dysbiosis
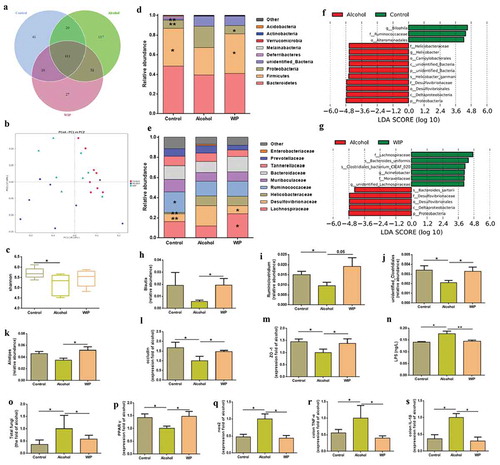
Figure 3. Identification of Meyerozyma guilliermondii as a casual fungus for AHS
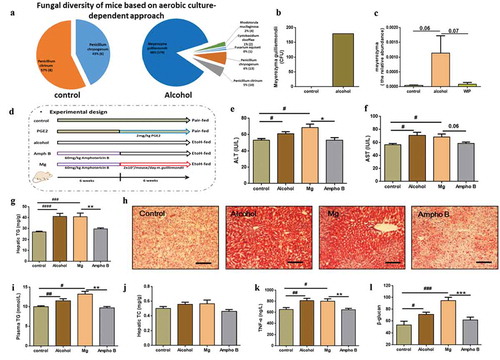
Figure 4. Contribution of fungi-induced PGE2 to alcoholic hepatic steatosis