Figures & data
Table 1. Differential expression patterns of 9 selected miRNAs in mouse BMDMs with or without exposure to C. albicans heat-killed yeasts or hyphae.
Figure 1. Expression levels of miRNAs in BMDMs exposed to UV-killed C. albicans cells. Expression levels of miR132 (A), miR146a (B), and miR155 (C) in C57Bl/6 BMDMs incubated with C. albicans UV-killed yeasts or hyphae at a MOI of 5. Expression levels were measured by qRT-PCR. Results represent the mean of fold change from 3 independent pooled experiments ± SD. Asterisks indicate p-value; (*) p < 0.05, (**) p < 0.01, (***) p < 0.001, (****) p < 0.0001.
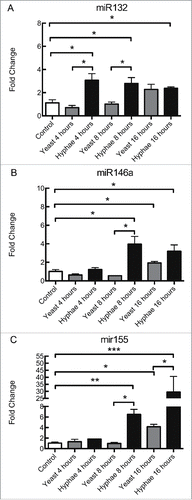
Figure 2. Dectin-1 is responsible for regulating miR155 expression in BMDMs exposed to C. albicans hypha. Expression levels of miR155 in (A) BMDMs obtained from different strains of C57Bl/6 mice, incubated for 8 hours with C. albicans UV-killed hyphae or in (B) Wild-type BMDMs incubated with either β-glucan or C. albicans UV-killed hyphae. (A) BMDMs from Wild-type mice and mice deficient in the PRRs TLR2, TLR4, both TLR2 and TLR4, or Dectin 1 (Wild-type, ΔTLR2, ΔTLR4, ΔTLR2/4 and ΔDectin1) were incubated for 8 hours with UV-killed C. albicans hyphae at a MOI of 5. (B) BMDMs from Wild-type C57Bl/6 mice were incubated for 8 hours with either β-glucan (depleted Zymosan) at 100 µg/mL or UV-killed C. albicans hyphae at a MOI of 5. Expression levels were measured by qRT-PCR. Results represent the mean of fold change from 3 independent pooled experiments ± SD. Asterisks indicate p-value; (*) p < 0.05, (**) p < 0.01, (***) p < 0.001, (****) p < 0.0001.
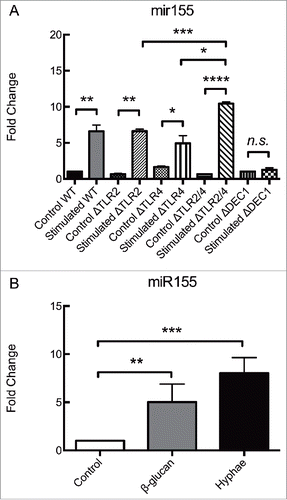
Figure 3. Dectin-1 activation of miR155 occurs via a Syk-dependent pathway. Effect of Dectin-1 pathway inhibitors on the expression of miR155 by different C57Bl/6 BMDMs with or without exposure to C. albicans UV-killed hyphae at an MOI of 5 for 8 hours. BMDMs with or without exposure to Dectin-1 inhibitors GW5074 (inhibits Raf-1), R406 (Syk inhibitor) or both compounds. Expression levels were measured by qRT-PCR. Results represent the mean of fold change from 3 independent pooled experiments ± SD. Asterisks indicate p-value; (*) p < 0.05, (**) p < 0.01, (***) p < 0.001, (****) p < 0.0001.
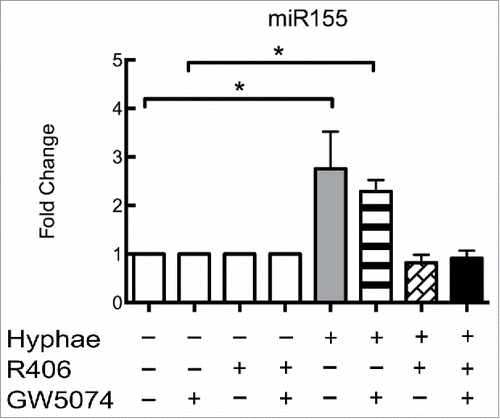
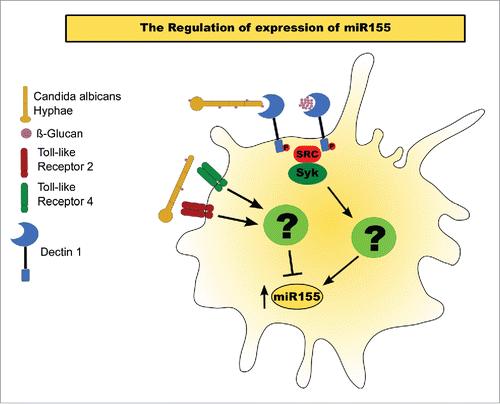
Figure 4. Model of miR155 activation in BMDMs exposed to C. albicans hyphae. miR155 activation by C. albicans. Exposure of β-glucans in the fungal cell wall interact with Dectin 1, inducing a Syk-dependent pathway that results in the upregulation of mature miR155. Activation of either TLR 2 or 4 inhibits the accumulation of the mature miRNA.