Figures & data
Figure 1. Schematics showing promising approaches to inhibiting oncogenic K-Ras: K-Ras activity can be inhibited directly or by agents that facilitate its dissociation from the plasma member. PDEδ binds to the PM-dissociated K-Ras and unloads it in the perinuclear region, whence K-Ras is translocated to the PtdSer- and SM-enriched recycling endosome (RE) for redelivery to the PM by vesicular transport. Disrupting its interaction with PDEδ or RE reduces the concentration of K-Ras at the PM. Also, perturbation of SM/Cer metabolism and distribution, which regulates PM PtdSer content, depletes PtdSer of the PM, resulting in K-Ras PM dissociation. Dysregulating RE activity by APEH inhibition further results in the mislocalization of PtdSer and K-Ras from the PM. PS – phosphatidylserine, SM – sphingomyelin, Cer – ceramide, ASM – acid sphingomyelinase, APEH – acylpeptide hydrolase, PDEδ – phosphodiesterase δ, G01 – a synthetic small molecule inhibitor of APEH
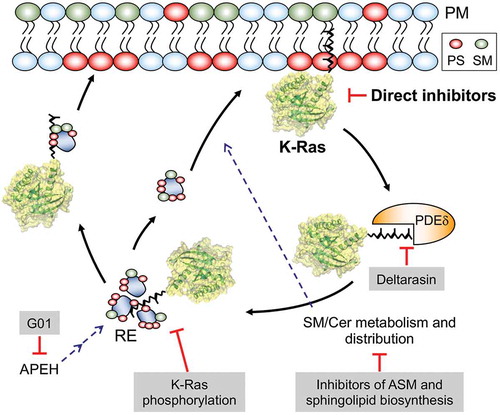
Figure 2. Druggable allosteric pockets on the catalytic domain of K-Ras. The sites most frequently targeted by published small molecules are pocket p1 near the core beta-sheet (pink) and pocket p2 between switch 2 and helix 3 (green). Pocket p3 near the C-terminus (blue) and pocket p4 behind switch 1 (yellow) are somewhat shallow and more polar than p1 and p2. See Grant et al. [Citation63], for a detailed structural analysis of the four pockets and Gupta et al. [Citation84], for a comparison of their druggability profile. Image reproduced from Grant et al. [Citation63]
![Figure 2. Druggable allosteric pockets on the catalytic domain of K-Ras. The sites most frequently targeted by published small molecules are pocket p1 near the core beta-sheet (pink) and pocket p2 between switch 2 and helix 3 (green). Pocket p3 near the C-terminus (blue) and pocket p4 behind switch 1 (yellow) are somewhat shallow and more polar than p1 and p2. See Grant et al. [Citation63], for a detailed structural analysis of the four pockets and Gupta et al. [Citation84], for a comparison of their druggability profile. Image reproduced from Grant et al. [Citation63]](/cms/asset/3f0cf1d8-c8d5-4cdb-aae6-b78343e7e2f9/ksgt_a_1655883_f0002_oc.jpg)
