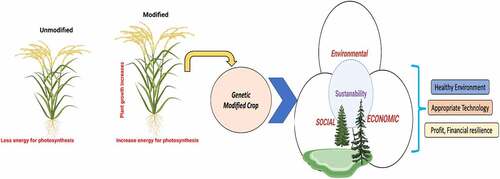
Figures & data
Table 1. Genetic modification application for abiotic stress
Table 2. Genetic modification applications for disease tolerance
Table 3. Genetic modification applications for nutrients