Figures & data
Figure 1. The photograph of Polypedilum henicurum. Scale bar indicates 500 μm. Photo credits: C. Song.

Figure 2. Mitochondrial genome map of Polypedilum henicurum. The CDS, tRNAs, rRNAs, GC skew and GC content are shown in the map.
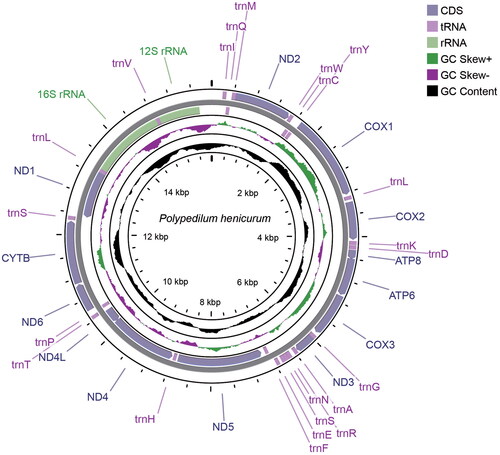
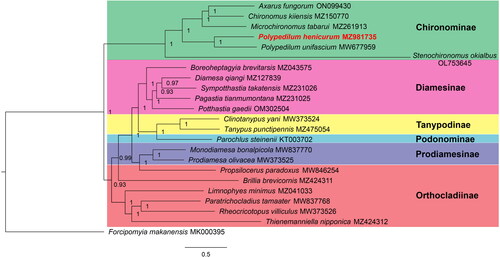
Figure 3. Bayesian inference phylogenetic tree of Chironomidae species. The tree was inferred from 22 registered Chironomidae species and one Ceratopogonidae species based on the concatenated sequences of mitogenomic PCGs. The following sequences were used: ON099430 (Qi et al. Citation2022), MZ150770 (Liu et al. Citation2022), MZ261913 (Kong et al. Citation2021), MZ981735 (this study), MW677959 (Lei et al. Citation2021), MZ043575 (Lin, Liu, et al. Citation2022), MZ127839 (Lin, Liu, et al. Citation2022), MZ231026 (Lin, Liu, et al. Citation2022), MZ231025 (Lin, Liu, et al. Citation2022), OM302504 (Lin, Liu, et al. Citation2022), MW373524 (Zheng et al. Citation2021), MZ475054 (Jiang et al. Citation2022), KT003702 (Kim et al. Citation2016), MW837770 (Lin, Zhao, et al. Citation2022), MW373525 (Zheng et al. Citation2021), MW846254 (Lin, Zhao, et al. Citation2022), MZ424311 (Lin, Zhao, et al. Citation2022), MZ041033 (Fang et al. Citation2022), MW837768 (Lin, Zhao, et al. Citation2022), MW373526 (Zheng et al. Citation2021), MZ424312 (Lin, Zhao, et al. Citation2022) and MK000395 (Jiang et al. Citation2019). Substitution model was GTR + I + G. BI posterior probability values are indicated at the nodes.
Supplemental Material
Download JPEG Image (971.8 KB)Data availability statement
The complete mitochondrial genome of Polypedilum henicurum for this study is openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov with accession number MZ981735. The associated BioProject, BioSample, and SRA are deposited in the Genome Sequence Archive in National Genomics Data Center at https://ngdc.cncb.ac.cn under accession numbers PRJCA015227, SAMC1125797, and CRA010003.
