Figures & data
Table 1 Isolation and purification details of secondary metabolite compounds, diameter of inhibition zone, MIC values (µg/mL), and MBC values (µg/mL) of active fractions against MRSA ATCC 43300
Figure 1 Neighbor-joining tree showing the relationship of strain SUK 25 based on a 16S rRNA gene sequences (1,450 nucleotides with closely related members of the genus Streptomyces omiyaensis NBRC 13449T and Kitasatospora setae KM-6054 as the outgroup).
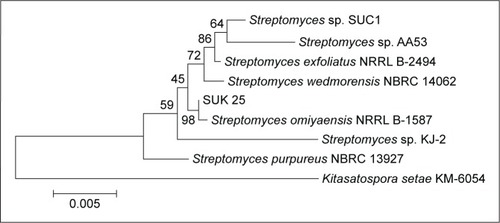
Figure 2 TLC analysis of whole-cell hydrolysate of SUK 25.
Abbreviations: TLC, thin-layer chromatography; SUK 25, strain Universiti Kebangsaan 25; DAP, diaminopimelic acid.
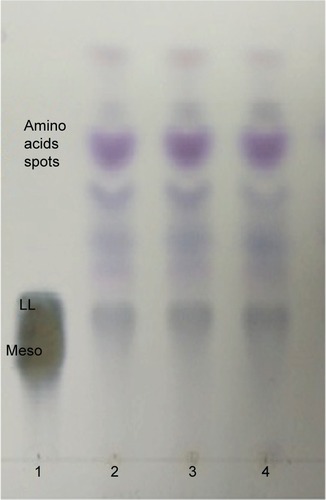
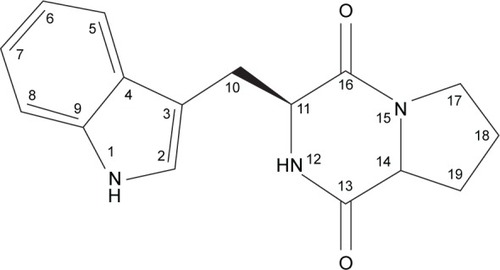
Table 2 1D-1H NMR and 13C NMR spectrum data of FVII-F2 cyclo-(l-tryptophanyl-l-prolyl)
Table 3 1D-1H NMR and 13C NMR spectrum data of FVII-F3 chloramphenicol
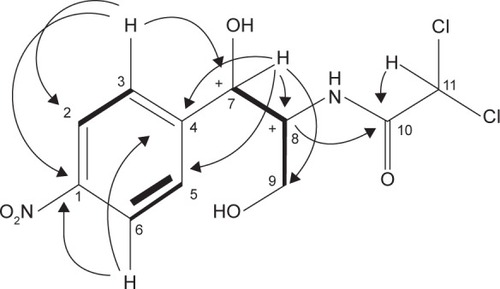
Figure 4 The chemical structure elucidation of chloramphenicol using 2D-NMR HMBC and COSY correlations of FVII-F3 compound.
Abbreviations: 2D-NMR, two-dimensional nuclear magnetic resonance; HMBC, heteronuclear multiple-bond correlation; COSY, correlations spectroscopy.