Figures & data
Table 1. Blood parameters.
Figure 2. (A) Representative hematoxylin and eosin (H&E) staining for the thoracic aortas (400× magnification). (B) Quantification of lesion area. (C) mRNA quantification for the antioxidant genes in aorta. Data are expressed as mean ± SEM (for each group, n = 10). ##P < 0.01 compared with control (CTL) rats; *P < 0.05 and **P < 0.01 compared with atherosclerotic (As) rats.
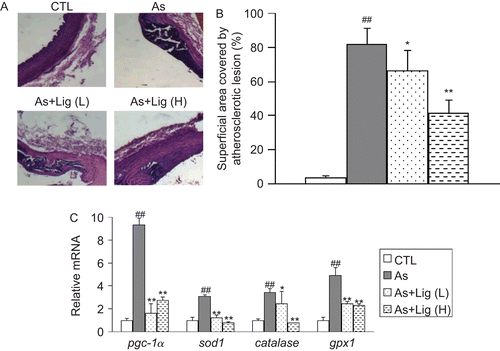
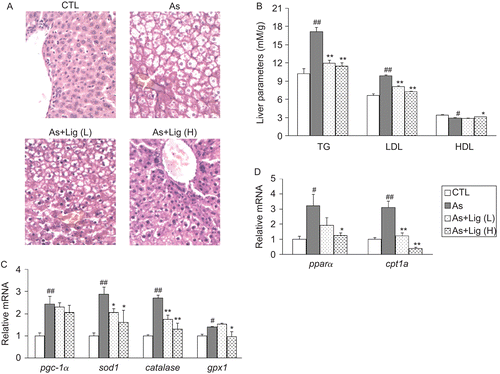
Figure 3. (A) Representative hematoxylin and eosin (H&E) staining for the livers (400× magnification). (B) Lipid profile in the livers. (C) mRNA quantification for the antioxidant genes. (D) mRNA quantification for the genes in fatty acid oxidation. Data are expressed as mean ± SEM (for each group, n = 10). #P < 0.05 and ##P < 0.01 compared with control (CTL) rats; *P < 0.05 and **P < 0.01 compared with atherosclerotic (As) rats.