Figures & data
Table 1. Crystal data and structure refinement details for compounds 1 and 2.
Figure 2. The molecular structure and atomic labeling scheme of 1. Displacement ellipsoids are drawn at the 40% probability level.
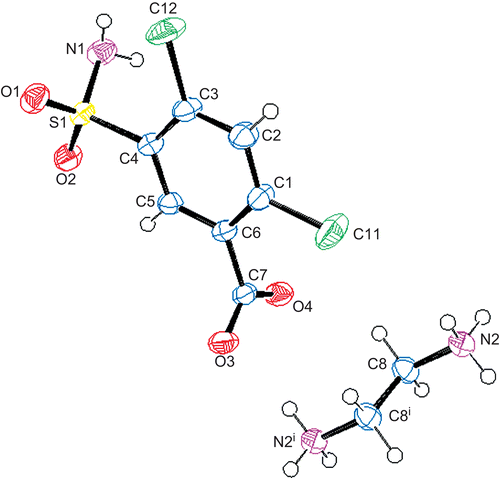
Figure 3. The molecular structure and atomic labeling scheme of 2. Displacement ellipsoids are drawn at the 40% probability level.
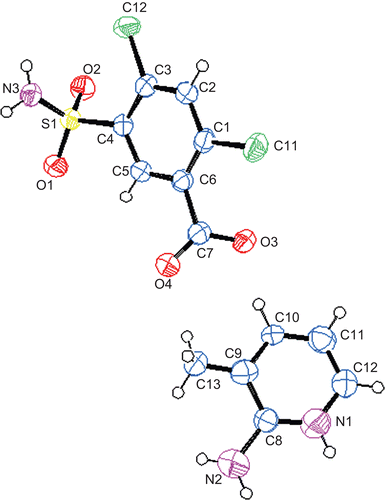
Figure 4. SDS-PAGE analyses of CA isoenzymes: (a) Standard hCA I, (b) isolated hCA I, (c) Standard hCA II and (d) isolated hCA II.
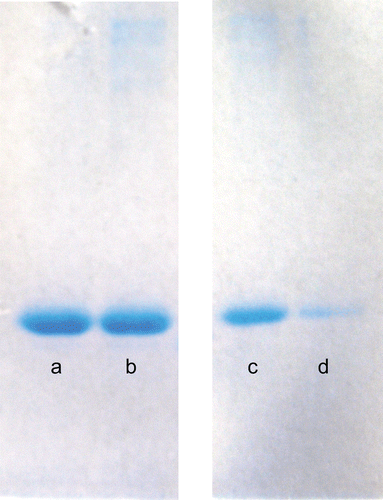
Table 2. Selected bond distances (Å) and angles (°) for compounds 1 and 2.
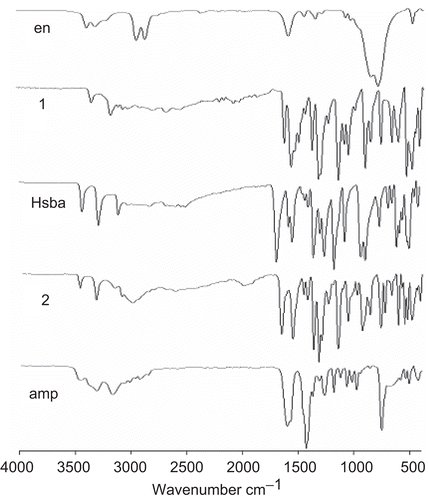
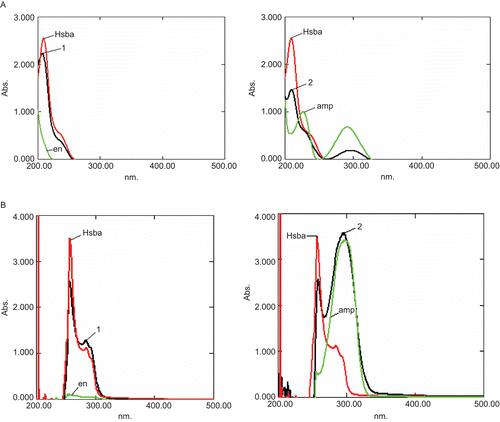
Table 3. Optical properties of compounds 1 and 2, and the free ligands amp and Hsba, in water and DMSO.
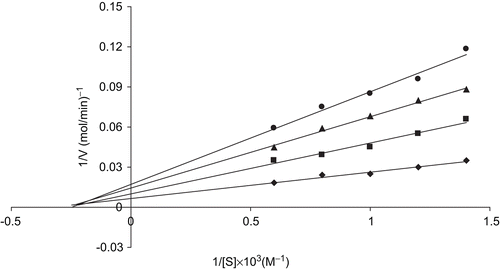
Table 4. IC50 and Ki values of hCA-I and hCA-II isoenzyme hydratase and esterase activities after inhibition experiments.