Figures & data
Figure 1. Chemical structures of the initial polyphenols (rutin 1, phloridzin 2 and esculin 3) with depicted positions of acylation.
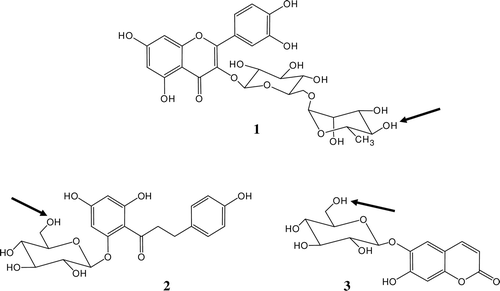
Table 1. Physicochemical parameters and conversion yields of rutin, phloridzin and esculin esters.
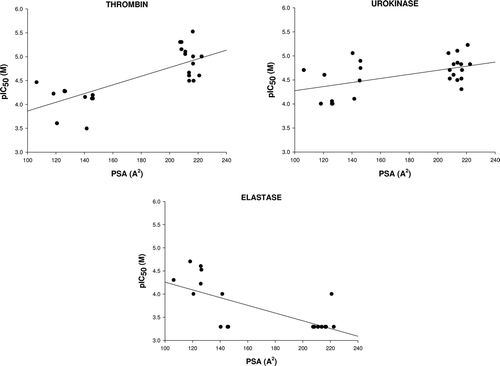
Table 2. Inhibitory activities of rutin, phloridzin, esculin and their lipophilic esters on trypsin, thrombin, urokinase and elastase expressed as concentrations required for 50% activity inhibition (IC50) (SD < 5%, n = 4, R > 0,95) and subsistent logarithmic values (–logIC50 +6)
Figure 2. Geometry of rutin and rutin stearate together with molecular lipophilic potential MLP (upper part) and molecular electrostatic potential MEP (lower). Potentials were calculated by VEGA program.
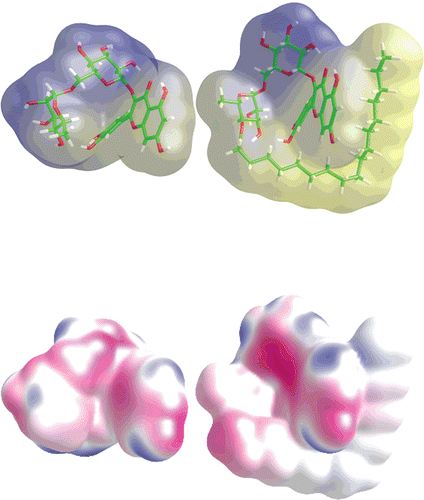
Table 3. Values of octanol–water partition coefficient AlogP for pH = 7.0, hydration energy Eaq, energy of the highest occupied molecular orbital EHOMO, energy of the lowest unoccupied molecular orbital ELUMO, dipole d, molecular volume V, molecular surface S, polar surface area PSA, number of hydrogen bond donors HBD and number of hydrogen bond acceptors HBA.
Table 4. Values of correlation coefficients obtained for simple linear regression for experimental inhibition activities for trypsin (TRP), urokinase (URK), thrombin (TRB) and elastase (ELS) and independent variables AlogP, Eaq, EHOMO, ELUMO, d, V, PSA, HBD and HBA.