Figures & data
Table 1. Summary of the statistical parameters obtained from the PLS analysis of five best CPHs are listed below.
Figure 1. Distance between the pharmacophore features of CPH1. The features like H-bond donor & electron withdrawing group and the aromatic rings are denoted as D, A and R, respectively.
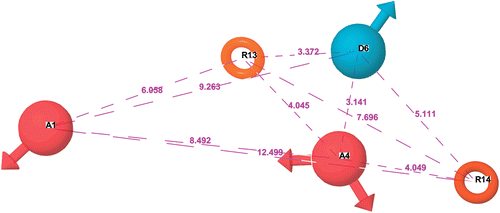
Figure 2. Visualization of 3D contours generated using CPH1 for the most active and inactive compounds (51 and 75). The subsets A and B depict the mapping of CPH1 onto the compounds 51 and 75, respectively. Subsets C-H depict the favorable (blue) and unfavorable (red) regions derived from the atom based 3D QSAR model for the H-bond donor (C, D), electron withdrawing (E, F) and hydrophobic properties (G, H) respectively.
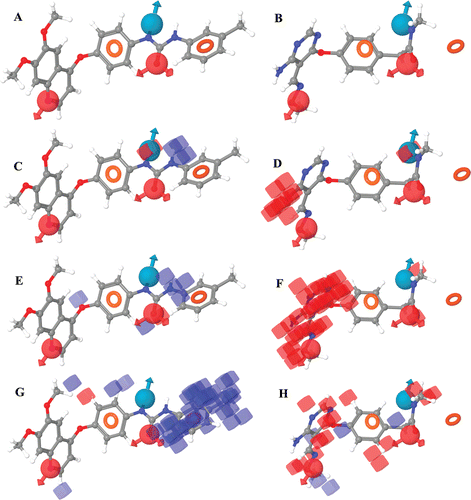
Figure 3. Plot of experimental activities against predicted biological activities of VEGFR-2 inhibitors. The traning and test set compounds are in spheres and diamonds, respectively.
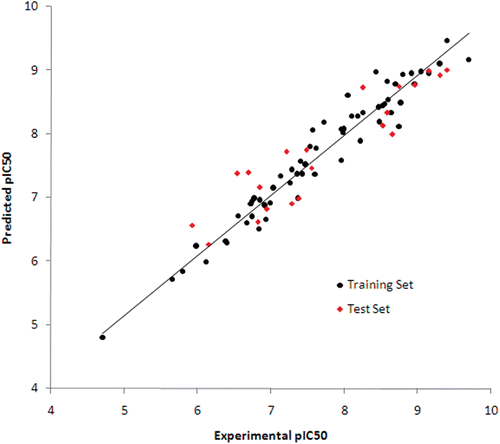
Figure 4. 2D representation of the interaction pattern of VEGFR-2 with active (A, B) and inactive (C, D) compounds before (A,C) and after (B,D) a simulation of 7 ns.
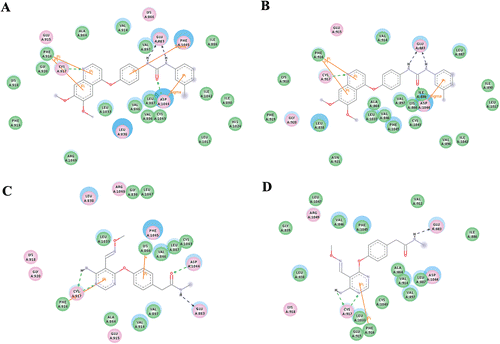
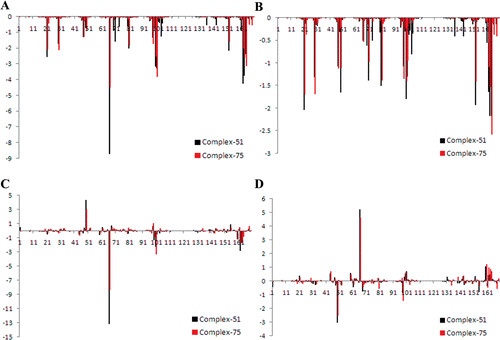
Figure 5. The RMSD (A) and RMSF (B) of a 7 ns long MD trajectory of VEGFR-2 complexes with active (black lines) and inactive (red lines) compounds, respectively.
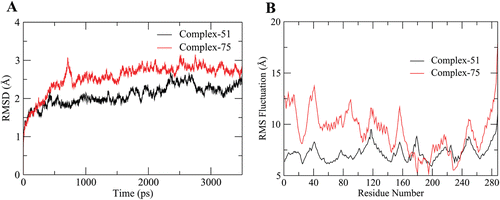
Table 2. Binding free energy and its individual components in kcal/mol calculated by the MMGBSA method.