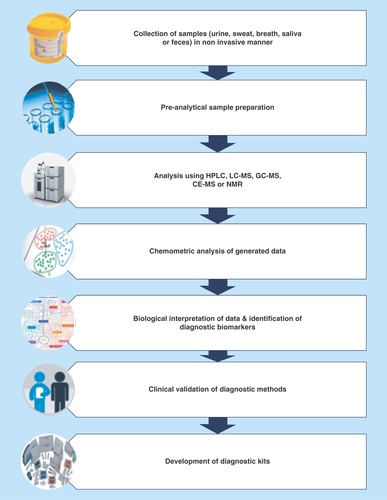
Figures & data
Table 1. Urinary metabolic profiling studies used for noninvasive diagnosis of different types of cancer.
Table 2. Urinary metabolic profiling studies used for noninvasive diagnosis of metabolic disorders and in born errors of metabolism.
Table 3. Urinary metabolic profiling studies used for noninvasive diagnosis of miscellaneous diseases and disorders.
Table 4. Salivary metabolic profiling studies used for noninvasive diagnosis of different diseases and disorders.
Table 5. Metabolic profiling studies using exhaled breath for noninvasive diagnosis of different diseases and disorders.
Table 6. Metabolic profiling studies using feces for noninvasive diagnosis of different diseases and disorders.