Figures & data
Table 1. Rfam classification of verified sRNAs from Xanthomonadaceae.
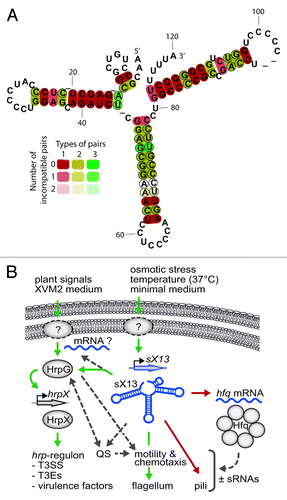
Figure 1. (A) Conservation of sX13. LocARNA prediction of the consensus secondary structure of all sX13 family members in xanthomonads listed in the Rfam database (Rfam 11.0, Aug. 2012).Citation29,Citation55 Compensatory mutations that retain the sX13 structure are indicated by circles. (B) Model of potential physiological functions of sX13 in Xcv according to Schmidtke et al.Citation24 sX13 accumulates in response to extracellular stimuli and indirectly promotes HrpG activation, which is induced by unknown signaling pathways.Citation56,Citation57 sX13 supposedly impacts on motility, quorum sensing (QS), and the activity of other sRNAs by repressing Hfq synthesis. Boxed arrows, wavy lines, and circles indicate genes, mRNAs, and proteins. Dashed circles mark unknown proteins or signaling pathways. Green, red, and dashed arrows denote positive, negative, and putative effects.