ABSTRACT
Pseudomonas aeruginosa is a major public health concern all around the world. In the frame of this work, a set of diverse environmental P. aeruginosa isolates with various antibiotic resistance profiles were examined in a Galleria mellonella virulence model. Motility, serotypes, virulence factors and biofilm-forming ability were also examined. Molecular types were determined by pulsed-field gel electrophoresis (PFGE). Based on our results, the majority of environmental isolates were virulent in the G. mellonella test and twitching showed a positive correlation with mortality. Resistance against several antibiotic agents such as Imipenem correlated with a lower virulence in the applied G. mellonella model. PFGE revealed that five examined environmental isolates were closely related to clinically detected pulsed-field types. Our study demonstrated that industrial wastewater effluents, composts, and hydrocarbon-contaminated sites should be considered as hot spots of high-risk clones of P. aeruginosa.
Introduction
Opportunistic Pseudomonas aeruginosa is a pathogen in diverse hosts including plants, animals and humans (Jander et al. Citation2000). In humans, it is responsible for severe nosocomial infections with the highest fatality rate of all Gram-negative bacteria (Aliaga et al. Citation2002). P. aeruginosa infections are multifactorial, the clinical outcome strongly depends on the interplay of cell-associated and extracellular virulence factors (Strateva and Mitov Citation2011; El Zowalaty et al. Citation2015). Intrinsic and acquired resistance to antibiotics is a co-factor in the mortality of P. aeruginosa infections (Cosgrove and Carmeli Citation2003). Swimming, swarming and twitching, the known motility forms of P. aeruginosa (Bradley Citation1980; Köhler et al. Citation2000), play a critical role in invasiveness (Drake and Montie Citation1988). Biofilms are well-documented as the source of infection (Kaiser et al. Citation2017), cause an increase in virulence (Schroeder et al. Citation2017) and are considered as hot spots for the accumulation and transfer of antibiotic resistance genes (ARGs) (Balcázar et al. Citation2015).
Due to its outstanding metabolic plasticity, P. aeruginosa can survive in diverse ecological niches including soil and aquatic habitats (Cabot et al. Citation2016), but it was usually observed in low abundance within these environments (Chatterjee et al. Citation2017). In contrast, P. aeruginosa was frequently isolated from hydrocarbon-contaminated sites (Kaszab et al. Citation2010) and can be dominant among hydrocarbon-degrading bacteria (Chaerun et al. Citation2004). The role of non-clinical environments as a reservoir or a transient recipient of P. aeruginosa is still under debate (Deredjian et al. Citation2014). Based on the absence of several virulence genes (Divya et al. Citation2018), or their non-haemolytic phenotype (Radhapriya et al. Citation2015), environmental P. aeruginosa isolates are sometimes considered as ‘non-pathogenic’ and are recommended for bioremediation (Ebadi et al. Citation2017), or plant protection purposes (Yasmin et al. Citation2017). Virulence models could provide tools for the reliable assessment of the actual virulence of these strains such as wax moth (Galleria mellonella), a non-mammalian test organism, introduced as an in vivo model for the study of P. aeruginosa (Velikova et al. Citation2016). Clinical P. aeruginosa is highly virulent to G. mellonella (Hill et al. Citation2014; Tsai et al. Citation2016) and the positive correlation between virulence of P. aeruginosa mutants in murine and G. mellonella models was verified (Jander et al. Citation2000). As the solo application of a non-mammalian model have its own limitations (Dubern et al. Citation2015), the simultaneous detection of virulence determinants, such as the presence of exotoxin and exoenzyme encoding genes or haemolytic activity, is proposed. Therefore, the overall aim of this research was to reach complex data on a set of environmental P. aeruginosa isolates with various antibiotic resistance profiles. To reach this goal, a virulence assay was performed in a G. mellonella model. Motility, serotype, virulence-related genes, haemolytic activity and biofilm-forming ability were also determined. To access genetic diversity, macrorestriction fragment patterns were created by PFGE.
Materials and methods
Bacteria isolation, identification, and screening for antibiotic resistance
Environmental samples were obtained from various environmental sources such as hydrocarbon (total petroleum hydrocarbons and/or benzene, toluene, ethylbenzene, xylene) contaminated groundwater and soil, compost, industrial wastewater effluents of Hungarian oil refineries and municipal sewage. P. aeruginosa isolates were cultivated by bacteriological standard methods and were identified with the PCR for the detection of species-specific variable regions V2 and V8 of 16S rDNA (Spilker et al. Citation2004). Antibiotic resistance profiles were determined with preliminary disc diffusion tests for 31 agents of 9 classes and quantitative Etest® (AB Biodisk, Solna, Sweden) for 9 chosen agents from the recommendation lists of the European Centre for Disease Prevention and Control (ECDC) and the Clinical Laboratory Standards Institute (CLSI) (EUCAST, Citation2018; CLSI Citation2018). Comparative isolates were P. aeruginosa ATCC-27853 (American Type Culture Collection, USA) and KPS3 (National Institute of Environmental Health, Hungary).
Serological typing
Serotypes of P. aeruginosa isolates were determined by agglutination tests using serotype-specific polyvalent antisera in accordance with the manufacturer’s instructions (Bio-Rad, France).
Pulsed-field gel electrophoresis (PFGE)
For PFGE, a slightly modified method of the previously described protocols (Bannerman et al. Citation1995; Deplano et al. Citation2005) was used with speI rare cutting enzyme for digestion and CHEF DRII System (Bio-Rad) for the separation of fragments in 1% agarose gel. Electrophoresis was in a 0.5x concentration Tris-borate-EDTA buffer (TBE) for 25 h at 5.8 V, 14°C with the initial switch time of 10 sec and the final switch time of 45 sec. Gels for PFGE were stained in 500 ml aqueous solution containing 0.5 μg/mL final concentration of ethidium-bromide, were washed with distilled water for 30 min and were photographed under UV transillumination. PFGE-generated DNA profiles were analyzed by the Fingerprinting II version 3.0 Software (Bio-Rad) using 1.0% position tolerance and 1.0% optimization settings. The universal size standard was the Salmonella Braenderup (H9812) strain. The quantitative differences among the banding patterns were defined by the Dice coefficient. Unweighted pair group method with arithmetic mean (UPGMA) was used for cluster analysis of the PFGE patterns. Interpretation of chromosomal DNA restriction patterns based on the criteria of Tenover et al. (Citation1995) using the following categories: indistinguishable (homology higher than 95%), closely related (homology between 85-95%), possibly related (homology higher than 85%, but the criteria of Tenover were not fulfilled) and different (homology lower than 85%). PFGE results of environmental isolates were compared to the surveillance database of the National Center for Epidemiology, Hungary on hospital outbreaks.
Haemolytic activity and virulence genes
Haemolytic activity was determined on Columbia agar with 5% sheep blood (Chemium Ltd., Hungary) with incubation parameters recommended by the manufacturer (37°C, 22 h). Microbial DNA was isolated by UltraClean Microbial DNA isolation kit (MoBio), following the manufacturer’s instructions. Virulence genes (effector protein-encoding genes exoS and exoU, elastase producing lasB, alginate encoding algD, alkaline phosphatase producing aprA and haemolytic phospholipase encoding plcH) were amplified using PCR with reaction parameters and primers described by Ullah et al. (Citation2017), Badamchi et al. (Citation2017) and Ajayi et al. (Citation2003). Results were visualized by agarose gel electrophoresis. Positive controls were P. aeruginosa ATCC 27853 for exoS, lasB, algD, aprA and P. aeruginosa KPS-3 (a nosocomial strain isolated in Hungary) for exoU and plcH genes.
Galleria mellonella infection model
G. mellonella test was performed as it was suggested by Koch et al. (Citation2014). Each test was performed in duplicate. Positive control was P. aeruginosa PA14 (DSM 19882), a virulent human clinical isolate, negative controls were injected with phosphate-buffered saline (PBS). After 24 and 48 h of incubation at 37°C, the rate of mortality was determined (a caterpillar was regarded as dead when no motion was observable). Virulence of the examined isolates was categorized after 48 h according to the survival of G. mellonella larvae into four groups: avirulent (with a survival rate of 75-100%), semi-virulent (survival rate: 50-74%), moderately virulent (survival rate: 25-49%) and virulent (survival rate: 0-24%).
Motility assays
Swimming, swarming and twitching were performed as it was previously described by Schmidt et al. (Citation2011). In brief, overnight bacterial cultures transferred to Luria-Bertani (LB) medium, were grown to exponential phase (OD600 1.5–2.5). Swimming was examined with Basal Medium 2 (BM2) containing 0.3% agar in 12 Well Nunc plates, inoculated with 0.5 µL liquid bacterial cultures. For swarming, BM2 was used with 0.5% agar content and an increased casamino acid concentration (0.5%) for a better flagellar synthesis. 1.0 µL of bacterial cultures were pipetted onto the prepared medium for inoculation. Twitching assay was performed on LB medium with 1% agar. Swimming and swarming plates were incubated overnight at 30°C, while the zone of twitching motility was marked after 48 h incubation at 37°C. The area was measured with the ImageJ program. The mean value of P. aeruginosa PA14 was calculated and was set as a threshold value for hypermotility. Categories and thresholds of motility tests are summarized in .
Table 1. Categorization of bacterial isolates according to their motility with threshold values
Biofilm and adherence assays
To assess biofilm-forming ability, a modified microtiter plate test was used with parameters and conditions described by Stepanović et al. (Citation2000) in TSB (Tryptic Soy Broth) medium. Test was performed in triplicate, isolates were classified after 24, 48, 72 and 96 h as no biofilm producers (0), weak biofilm producers (+), moderate biofilm producers (++), or strong biofilm producers (+++). The cut-off OD (ODc) was determined as it was recommended by Stepanović et al. (Citation2000).
Biofilm production was visualized by confocal laser scanning microscopy (CLSM) in Twincore Centre (Hanover, Germany) to acquire images of 48 h biofilms grown in static 96-well plate assays. The method of visualization was described previously (Van Duuren et al. Citation2017).
Results of environmental P. aeruginosa were challenged by a pairwise statistical analysis to clarify the possible relations among different features. Pearson’s correlation (r) was calculated with a confidence interval of 95%.
Results
Antibiotic susceptibility
Based on their antibiotic resistance profiles, 44 non-duplicate environmental P. aeruginosa isolates were chosen for further investigations representing sensitive (n = 29) and resistant (n = 15) phenotypes. Resistant isolates were detectable in compost and hydrocarbon-contaminated groundwater or soil. Resistance was widespread against Cefotaxime (18.2%), Ceftriaxone (25.0%) and Imipenem (25.0%), while Cefepime, Ciprofloxacin and Gentamicin remained effective against nearly all examined isolates. Four environmental strains (P14, P43, P46 and P69) met the MDR (multidrug resistance) criteria defined by Magiorakos et al. (Citation2012). Strains isolated from oil refinery sewage showed a sensitive phenotype. The origin and the antibiotic resistance profiles of environmental isolates are shown in .
Table 2. The origin and antibiotic resistance profiles of environmental P. aeruginosa isolates
Serotypes and PFGE patterns
Among environmental isolates, nine serotypes were determined with the dominance of O6 (29.5%), O1 (15.9%) and O3 (15.9%). The remaining 17 isolates were typed as O1, O4, O7, O9, O9, or were non-typable (ONT). Based on their PFGE banding patterns, 18 (40.9%) environmental isolates were considered as sporadic strains with a loose relationship to other environmental or nosocomial profiles (homology was lower than 85%). Three environmental isolates (P16, P80 and P174) showed 84-88% homology in PFGE-profiles with clinical isolates (originating from hospitals located in Budapest), and five of the examined 44 environmental isolates, (P59, P119, P124, P125 and P172) showed the same PFGE-types as a previously described nosocomial strain (with 85-95% homology). The common origin of these five environmental strains with isolates of hospital outbreaks is presumed. At the same time, none of the examined environmental isolates had 95% or higher homology with the evaluated nosocomial fingerprints (.).
Figure 1. The characteristics of environmental P. aeruginosa isolates in order of their localization on PFGE dendrogram. grey coloured, same pulsed-field types as clinical counterparts; bold font, multidrug resistance (MDR); nd, no data; *clinical reference strains; ** + positive PCR; – negative PCR; ***- no haemolysis; + moderate haemolysis, ++ normal haemolysis, +++ intensive haemolysis; **** Biofilm-forming in modified microtiter assay: – no biofilm producer; + weak biofilm producer; ++ moderate biofilm producer; +++ strong biofilm producer
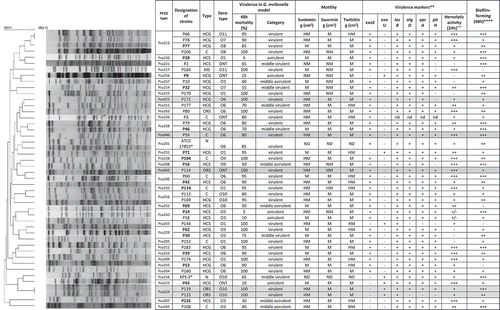
Environmental isolates showing 85-95% homology with clinical counterparts (marked with grey in .) were sensitive to the examined antibiotics (except Imipenem and Cefotaxime-resistant P16), had clinically relevant serotypes O6 and O10 (with the exception of non-typable strain P124) and had various virulence gene profiles. The shared features of these clinically related environmental isolates were as follows: motility or hypermotility in all examined forms of motion, virulence (80-100% mortality) in the G. mellonella model, haemolytic activity and biofilm-forming capability. Three of these environmental isolates originated from industrial sewage effluents.
Virulence genes
The frequency of the examined virulence-related genes among environmental P. aeruginosa was as follows: lasB (100.0%), algD (95.3%), aprA (95.3%), plcH (88.3%), exoS (72.7%) and exoU (22.7%). All environmental isolates harboured at least two of the examined six virulence-related genes and 34 (79.0%) isolates showed a profile with five simultaneously detectable virulence genes. Regarding exoU/exoS, the most variable traits among virulence genes (Ahmed et al. Citation2019), the commonly determined environmental profile is possessing exoS. However, environmental isolates carrying exoU (isolated from hydrocarbon-contaminated soil, groundwater and wastewater) and isolates lacking both exoU/exoS (isolated from compost samples) were also detected. Regarding haemolytic features, 15 isolates (34.1%) showed intensive beta-haemolytic activity and only seven (15.9%) of the environmental isolates were non-haemolytic (.).
Virulence in G. mellonella model
In the applied G. mellonella virulence model, the majority (65.9%) of the examined environmental isolates was virulent with a mortality rate of 75-100%. Five of the examined 44 environmental P. aeruginosa isolates were proved to be moderately virulent (50-75%), five isolates were semi-virulent (25-50%) and five strains were avirulent (0-25% mortality). The survival slope variations between virulent and not virulent (moderately, semi, or avirulent) phenotypes suggesting a significant difference in virulence after 24 h injection (.).
Figure 2. Galleria survival after inoculation with bacterial samples. Non-virulent environmental (48 h mortality between 0-75%); virulent environmental isolates (48 h mortality between 75-100%)
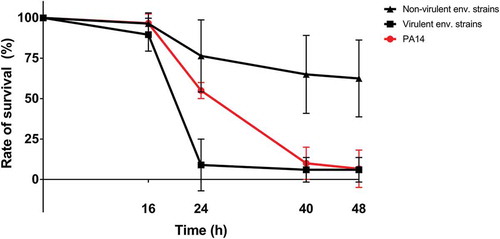
Regarding motility assay, all examined environmental isolates showed at least one type of motility and one strain (P. aeruginosa P114 originated from compost) was proved to be hypermotile in all examined motility forms (.). Twenty-five (56.8%) environmental isolates were hypermotile in swimming and 13 (29.5%) strains showed hypermotility in twitching. Swarming was moderate in frequency since only two (4.5%) isolates were proved to be hypermotile.
Biofilm-forming ability
Regarding biofilm-forming ability, different incubation times (24, 48, 72 and 96 h) lead to huge differences in the detected level of adherence. Therefore, the recommended 48 h incubation (Van Duuren et al. Citation2017) was used for further evaluation in the applied microtiter-plate test. After 48 h, only three (6.8%) of the examined environmental isolates were non-adherent, 61.3% was weak or moderate biofilm producer and 14 (31.8%) was proved to be a strong biofilm producer. CLSM revealed, that 14 (34.1%) of the biofilm-producing P. aeruginosa isolates (of which four were strongly adherent) did not form structured biofilms: the detected OD values were a result of the massive accumulation of planktonic cells in pellicles (.). Twenty-seven (61.3%) of the examined 44 environmental isolates showed biofilm-forming ability with both applied techniques of which 10 (22.7%) was a strong biofilm producer.
Figure 3. The biofilm-forming ability of environmental P. aeruginosa isolates P14 and P28 based on their adherence in microtiter-plate test and their image by confocal laser scanning microscopy (CLSM). *OD – optical density; **ODc – cut-off OD, threshold of weak-adherence; ***2xODc – threshold of moderate adherence; ****4xODc – the threshold of strong adherence
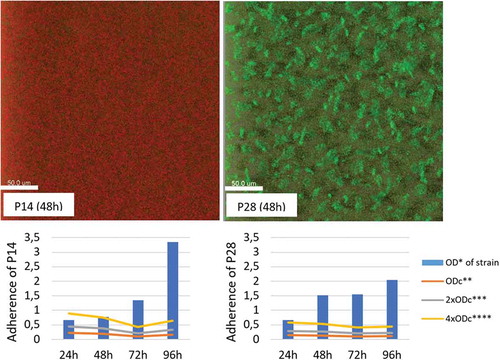
Statistical analysis
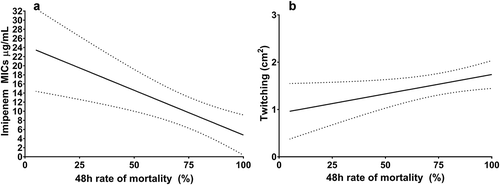
The statistical analysis revealed that Minimal Inhibitory Concentrations (MICs) against six different agents (Gentamicin, Imipenem, Cefepime, Ceftazidime, Cefotaxime and Cefoperazone-sulbactam) showed inverse correlation with the rate of 48 h mortality in G. mellonella test: the higher MICs (increasing resistance) lead to a lower rate of in vivo mortality (. part A). Despite the negative correlation, simultaneously virulent and antibiotic-resistant phenotypes were detectable among isolates originating from hydrocarbon-contaminated groundwater (P53, P62, P71, P77) soil (P16, P42) or compost (P114). The presence or absence of the examined virulence-related genes lasB, algD, aprA, plcH, exoS and exoU and the phenotypically determined haemolytic activity did not show significant correlation with G. mellonella mortality results. The only exception was the simultaneous lack of exoS and exoU genes showing a significantly lower rate of mortality in the applied virulence model. Twitching was a possible motility-related contributor in virulence on G. mellonella, since it was the only examined motility form showing a significant correlation with the rate of mortality (. part B). The cumulative ability for motion also significantly enhanced the rate of virulence, which underlines that the critical role of twitching in G. mellonella infection can be at least partially replaced by swimming and/or swarming. Hypermotility tends to be a lacking ability of avirulent environmental isolates with one exception (P14) only.
Discussion
According to the scientific literature, the detected rate of resistance against Piperacillin (13.6%) and especially Imipenem (25.0%) was higher than as it was recently reported by Liew et al. (Citation2019) on freshwater P. aeruginosa isolates. In the present study, Imipenem-resistant isolates were exclusively originated from hydrocarbon-contaminated samples verifying the importance of contaminated areas as a possible source of carbapenem-resistant cell lines (Kaszab et al. Citation2010). Regarding dominant serotypes, the detection rate of O6 (29.5%) was lower, than as it was previously reported in marine habitats (95.8%) (Bouhaddioui et al. Citation2002), which emphasize that groundwater, soil and compost can be more diverse in P. aeruginosa serotypes than marine environments. Clinically relevant serotypes determined by Lu et al. (Citation2014) were detected in 26 cases, meaning that 59.0% of the examined environmental isolates were in a clinically important serogroup (O1, O6, O10 or O11). At the same time, O12, the predominant MDR serotype in clinical settings (Thrane et al. Citation2015) was not detectable. G. mellonella virulence test revealed, that environmental strains with clinically relevant serotypes have a higher rate of virulent isolates (84.6%) than strains with non-clinical (O3, O4, O7, O8, O9 and ONT) serotypes (33.3%).
Based on our PFGE results, PFGE profiles did not show a significant correlation with virulence, biofilm or motility properties of the examined isolates: virulent and antibiotic-resistant strains were localized randomly on dendrogram. Although some studies previously reported a correlation between PFGE types and antibiotic resistance (Salimi et al. Citation2010), our findings agree with the reports of no correlation between these features (Tokajian et al. Citation2012; Karami et al. Citation2018). This result emphasizes the lack of discriminatory power of phenotypic methods and support the necessity of genomic studies in this field.
Among the examined virulence genes, the determined frequency of invasive exoS (72.7%) and cytotoxic exoU (22.7%) are within the recently observed ranges in the environmental (Rutherford et al. Citation2018; Ahmed et al. Citation2019) and clinical (Fazeli and Momtaz Citation2014) settings. Isolates harbouring both exoS and exoU virulence genes were not detectable, since these virulence genes are nearly always mutually exclusive (Shaver and Hauser Citation2004). The role of virulence genes and their effector proteins in G. mellonella killing was previously examined and it was concluded that only the secretion of ExoT and ExoU plays a significant role (Miyata et al. Citation2003). Here we’ve verified that exoS is also an important contributor in wax moth killing and only the simultaneous absence of exoS and exoU leading to a decreased rate of mortality. Considering the low number of isolates simultaneously lacking exoU and exoS, further investigations and gene expression analyses are required to verify these results.
Regarding extracellular protease encoding lasB (elastase LasB) and aprA (alkaline protease AprA), lasB was detected in a higher rate (100.0%) than in clinical settings (Roshani-Asl et al. Citation2018). Due to this exclusiveness, it was not possible to evaluate the previously reported relationship between the presence of lasB gene and biofilm-forming ability or Imipenem, Piperacillin/Tazobactam resistance (Roshani-Asl et al. Citation2018). Regarding aprA, the previously reported clinical frequencies were 27.2% among patients with otitis media (Hassuna Citation2016), 28.6% among burned patients (Fazeli and Momtaz Citation2014) and 100% in intensive care units (ICUs) (Bradbury et al. Citation2010). The rate of aprA detection (95.3%) among our environmental isolates is similar to its frequency among clinical strains isolated from ICUs. AlgD (97.7%) and plcH (86.6%) were both detected in the same values as in clinical environments (92.3–98.0% and 84.6%, respectively) (Taee et al. Citation2014; Georgescu et al. Citation2016). Interestingly, there was no significant correlation between the presence of plcH gene, encoding a haemolytic phospholipase C precursor, and the phenotypically determined haemolytic activity of environmental strains. The answer to this insignificant correlation may be the role of other cytolytic factors in haemolysis such as the production of exotoxin A or elastase (Lee et al. Citation2006).
Our G. mellonella assay verified that virulence is a dominant phenotypic trait in the environment. Virulent isolates with a notable set of invasive and cytotoxic factors were isolated from all examined environmental media (groundwater, soil, compost, sewage effluents). This finding supports the opinion that all P. aeruginosa strains are potential pathogens (Pirnay et al. Citation2009; Silby et al. Citation2011) due to their invasive and toxigenic features (Mesaros et al. Citation2007; Kumar et al. Citation2013). Considering that, due to host-specific virulence factors (Choi et al. Citation2002), virulence in G. mellonella model can significantly differ from other test organisms, additional virulence tests should be performed in the future.
Based on our motility results, the previously drawn connections between twitching and biofilm formation (O’Toole and Kolter Citation1998) or antibiotic resistance (Shehata and Sayed Citation2011) were not verified since twitching motility was widespread among environmental isolates and the limitations of microtiter-plate test for detecting biofilms were revealed.
Regarding biofilm-forming ability, the detected rate of strong biofilm producer environmental isolates (22.7%) is closer to the previously reported rate for human isolates (23.5%) than for strains isolated from animals (9%) (Milivojevic et al. Citation2018). The fact that 34.8% of the positive microtiter-plate results were not structured biofilms emphasize the importance of visual techniques in biofilm detection (Van Duuren et al. Citation2017).
Regarding the correlation between virulence and antibiotic resistance features, the connection was previously reported to be both positive (Neidig et al. Citation2013) and negative (Bartoli et al. Citation2015), but recent studies debated inverse correlation (Schroeder et al. Citation2017; Al Dawodeyah et al. Citation2018). Based on the results presented here, the negative relationship of antibiotic resistance and virulence is still possible in the environment.
Conclusions
A set of 44 environmental P. aeruginosa were examined isolated from various (e.g. hydrocarbon-contaminated) environmental sources. The majority of environmental P. aeruginosa was proved to be virulent with a verified ability for motility, haemolysis and biofilm-forming. Five examined environmental isolates were closely related to clinically detected PFGE types. The presence or absence of the revealed arsenal of virulence-related factors or genes did not correlate with phenotypic traits, which emphasize the complexity of virulence-related mechanisms. Despite the negative correlation between antibiotic resistance and wax moth killing, simultaneously resistant and virulent clones can be found among environmental isolates. The role of various environmental niches as a possible source of pathogen and resistant lineages is presumed: industrial wastewater effluents, composts and hydrocarbon-contaminated sites should be considered as hot spots of pathogen and resistant P. aeruginosa.
Conflict of interest
No potential conflict of interest was reported by the authors.
Acknowledgments
Authors gratefully acknowledge the support of Professor Dr Susanne Häußler, Helmholz Centre for Infection Research, Germany.
Additional information
Funding
References
- Ahmed ABH, Abbassi MS, Rojo-Bezares B, Ruiz-Roldán L, Dhahri R, Mehri I, Sáenz Y, Hassen A. 2019. Characterization of Pseudomonas aeruginosa isolated from various environmental niches: new STs and occurrence of antibiotic susceptible “high risk clones”. Int J Env Health Res. doi:https://doi.org/10.1080/09603123.2019.1616080.
- Ajayi T, Allmond LR, Sawa T, Wiener-Kronish JP. 2003. Single-nucleotide-polymorphism mapping of the Pseudomonas aeruginosa type III secretion toxins for development of a diagnostic multiplex PCR system. J Clin Microbiol. 41(8):3526–3531. doi:https://doi.org/10.1128/JCM.41.8.3526-3531.2003.
- Al Dawodeyah HY, Obeidat N, Abu-Qatouseh LF, Shehabi AA. 2018. Antimicrobial resistance and putative virulence genes of Pseudomonas aeruginosa isolates from patients with respiratory tract infection. Germs. 8(1):31–40. doi:https://doi.org/10.18683/germs.2018.1130.
- Aliaga L, Mediavilla JD, Cobo F. 2002. A clinical index predicting mortality with Pseudomonas aeruginosa bacteraemia. J Med Microbiol. 51(7):615–619. doi:https://doi.org/10.1099/0022-1317-51-7-615.
- Badamchi A, Masoumi H, Javadinia S, Asgarian R, Tabatabaee A. 2017. Molecular detection of six virulence genes in Pseudomonas aeruginosa isolates detected in children with urinary tract infection. Microb Pathog. 107:44–47. doi:https://doi.org/10.1016/j.micpath.2017.03.009.
- Balcázar JL, Subirats J, Borrego CM. 2015. The role of biofilms as environmental reservoirs of antibiotic resistance. Front Microbiol. 6:1216. doi:https://doi.org/10.3389/fmicb.2015.01216.
- Bannerman TL, Hancock GA, Tenover FC, Miller JM. 1995. Pulsed-field gel electrophoresis as a replacement for bacteriophage typing of Staphylococcus aureus. J Clin Microbiol. 33(3):551–555.
- Bartoli C, Lamichhane JR, Berge O, Varvaro L, Morris CE. 2015. Mutability in Pseudomonas viridiflava as a programmed balance between antibiotic resistance and pathogenicity. Mol Plant Pathol. 16(8):860–869. doi:https://doi.org/10.1111/mpp.12243.
- Bouhaddioui B, Ben Slama K, Gharbi S, Boudabous A. 2002. Epidemiology of clinical and environmental Pseudomonas aeruginosa strains. Ann Microbiol. 52(3):223–235.
- Bradbury RS, Roddam LF, Merritt A, Reid DW, Champion AC. 2010. Virulence gene distribution in clinical, nosocomial and environmental isolates of Pseudomonas aeruginosa. J Med Microbiol. 59:881–890. doi:https://doi.org/10.1099/jmm.0.018283-0.
- Bradley DE. 1980. A function of Pseudomonas aeruginosa PAO polar pili: twitching motility. Can J Microbiol. 26(2):146–154. doi:https://doi.org/10.1139/m80-022.
- Cabot G, Zamorano L, Moyà B, Juan C, Navas A, Blázquez J, Oliver A. 2016. Evolution of Pseudomonas aeruginosa antimicrobial resistance and fitness under low and high mutation rates. Antimicrob Agents Chemother. 60(3):1767–1778. doi:https://doi.org/10.1128/AAC.02676-15.
- Chaerun SK, Tazaki K, Asada R, Kogure K. 2004. Bioremediation of coastal areas 5 years after the Nakhodka oil spill in the Sea of Japan: isolation and characterization of hydrocarbon-degrading bacteria. Environ Int. 30(7):911–922. doi:https://doi.org/10.1016/j.envint.2004.02.007.
- Chatterjee P, Davis E, Yu F, James S, Wildschutte JH, Wiegmann DD, Sherman DH, Mckay RM, Lipuma JJ, Wildschutte H. 2017. Environmental Pseudomonas inhibit cystic fibrosis patient-derived Pseudomonas aeruginosa. Appl Environ Microbiol. 83:e02701–16. doi:https://doi.org/10.1128/AEM.02701-16.
- Choi JY, Sifri CD, Goumnerov BC, Rahme LG, Ausubel FM, Calderwood SB. 2002. Identification of virulence genes in a pathogenic strain of Pseudomonas aeruginosa by representational difference analysis. J Bacteriol. 184(4):952–961. doi:https://doi.org/10.1128/jb.184.4.952-961.2002.
- CLSI. 2018. Performance standards for antimicrobial susceptibility testing. 28th ed. Wayne (PA): Clinical and Laboratory Standards Institute. CLSI Supplement M100.
- Cosgrove SE, Carmeli Y. 2003. The impact of antimicrobial resistance on health and economic outcomes. Clin Infect Dis. 36:1433–1437. doi:https://doi.org/10.1086/375081.
- Deplano A, Denis O, Poirel L, Hocquet D, Nonhoff C, Byl B, Nordmann P, Vincent JL, Struelens MJ. 2005. Molecular characterization of an epidemic clone of panantibiotic-resistant Pseudomonas aeruginosa. J Clin Microbiol. 43(3):1198–1204. doi:https://doi.org/10.1128/JCM.43.3.1198-1204.2005.
- Deredjian A, Colinon C, Hien E, Brothier E, Youenou B, Cournoyer B, Dequiedt S, Hartmann A, Jolivet C, Houot S, et al. 2014. Low occurrence of Pseudomonas aeruginosa in agricultural soils with and without organic amendment. Front Cell Infect Microbiol. 4:53. doi:https://doi.org/10.3389/fcimb.2014.00053.
- Divya J, Mohandas A, Singh B. 2018. A non-pathogenic environmental isolate of Pseudomonas aeruginosa MCCB 123 with biotechnological potential. Int J Curr Microbiol Appl Sci. 7(1):3060–3071. doi:https://doi.org/10.20546/ijcmas.2018.701.363.
- Drake D, Montie TC. 1988. Flagella, motility and invasive virulence of Pseudomonas aeruginosa. J Gen Microbiol. 134(1):43–52. doi:https://doi.org/10.1099/00221287-134-1-43.
- Dubern JF, Cigana C, De Simone M, Lazenby J, Juhas M, Schwager S, Bianconi I, Döring G, Eberl L, Williams P, et al. 2015. Integrated whole-genome screening for Pseudomonas aeruginosa virulence genes using multiple disease models reveals that pathogenicity is host specific. Environ Microbiol. 17(11):4379–4393. doi:https://doi.org/10.1111/1462-2920.12863.
- Ebadi A, Khoshkholgh Sima NA, Olamaee M, Hashemi M, Ghorbani Nasrabadi R. 2017. Effective bioremediation of a petroleum-polluted saline soil by a surfactant-producing Pseudomonas aeruginosa consortium. J Adv Res. 8:627–633. doi:https://doi.org/10.1016/j.jare.2017.06.008.
- El Zowalaty ME, Al Thani AA, Webster TJ, El Zowalaty AE, Schweizer HP, Nasrallah GK, Marei HE, Ashour HM. 2015. Pseudomonas aeruginosa: arsenal of resistance mechanisms, decades of changing resistance profiles, and future antimicrobial therapies. Future Microbiol. 10(10):1683–1706. doi:https://doi.org/10.2217/fmb.15.48.
- European Committee on Antimicrobial Susceptibility Testing. 2018. Breakpoint tables for interpretation of MICs and zone diameters–version 9.0. [accessed 2018 July 19]. http://www.eucast.org.
- Fazeli N, Momtaz H. 2014. Virulence gene profiles of multidrug-resistant Pseudomonas aeruginosa isolated from Iranian hospital infections. Iran Red Crescent Med J. 16(10):e15722. doi:https://doi.org/10.5812/ircmj.61469.
- Georgescu M, Gehorghe I, Curutiu C, Lazar V, Bleotu C, Chifiriuc M-C. 2016. Virulence and resistance features of Pseudomonas aeruginosa strains isolated from chronic leg ulcers. BMC Infect Dis. 16(Suppl 1):92. doi:https://doi.org/10.1186/s12879-016-1396-3.
- Hassuna NA. 2016. Molecular detection of the virulent exoU genotype of Pseudomonas aeruginosa isolated from infected surgical incisions. Surg Inf. 17(5):610–614. doi:https://doi.org/10.1089/sur.2016.065.
- Hill L, Veli N, Coote PJ. 2014. Evaluation of Galleria mellonella larvae for measuring the efficacy and pharmacokinetics of antibiotic therapies against Pseudomonas aeruginosa infection. Int J Antimicrob Agents. 43(3):254–261. doi:https://doi.org/10.1016/j.ijantimicag.2013.11.001.
- Jander G, Rahme LG, Ausubel FM. 2000. Positive correlation between virulence of Pseudomonas aeruginosa mutants in mice and insects. J Bacteriol. 182(13):3843–3845. doi:https://doi.org/10.1128/JB.182.13.3843-3845.2000.
- Kaiser SJ, Mutters NT, DeRosa A, Ewers C, Frank U, Günther F. 2017. Determinants for persistence of Pseudomonas aeruginosa in hospitals: interplay between resistance, virulence and biofilm formation. Eur J Clin Microbiol Infect Dis. 36(2):243–253. doi:https://doi.org/10.1007/s10096-016-2792-8.
- Karami P, Mohajeri P, Mashouf RY, Karami M, Yaghoobi MH, Dastan D, Alikhani MY. 2018. Molecular characterization of clinical and environmental Pseudomonas aeruginosa isolated in a burn center. Saudi J Biol Sci. doi:https://doi.org/10.1016/j.sjbs.2018.07.009.
- Kaszab E, Kriszt B, Atzél B, Szabó G, Szabó I, Harkai P, Szoboszlay S. 2010. The occurrence of multidrug-resistant Pseudomonas aeruginosa on hydrocarbon-contaminated sites. Microb Ecol. 59(1):37–45. doi:https://doi.org/10.1007/s00248-009-9551-7.
- Koch G, Nadal-Jimenez P, Cool RH, Quax WJ. 2014. Assessing Pseudomonas virulence with nonmammalian host: Galleria mellonella. In: Filloux A, Ramos J-L, editors. Pseudomonas methods and protocols, methods in molecular biology. Vol. 1149. Springer Science+Business Media New York. doi:https://doi.org/10.1007/978-1-4939-0473-0_52.
- Köhler T, Curty LK, Barja F, Van Delden C, Pechere JC. 2000. Swarming of Pseudomonas aeruginosa is dependent on cell-to-cell signaling and requires flagella and pili. J Bacteriol. 182(21):5990–5996. doi:https://doi.org/10.1128/JB.182.21.5990-5996.2000.
- Kumar A, Munder A, Aravind R, Eapen SJ, Tümmler B, Raaijmakers JM. 2013. Friend or foe: genetic and functional characterization of plant endophytic Pseudomonas aeruginosa. Environ Microbiol. 15(3):764–779. doi:https://doi.org/10.1111/1462-2920.12031.
- Lee DG, Urbach JM, Wu G, Liberati NT, Feinbaum RL, Miyata S, Diggins LT, He J, Saucier M, Déziel E, et al. 2006. Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol. 7(10):R90. doi:https://doi.org/10.1186/gb-2006-7-10-r90.
- Liew SM, Rajaskaram G, Puthucheary SDA, Chua KH. 2019. Antimicrobial susceptibility and virulence genes of clinical and environmental isolates of Pseudomonas aeruginosa. Peer J. 7e:6217. doi:https://doi.org/10.7717/peerj.6217.
- Lu Q, Eggimann P, Luyt C-E, Wolff M, Tamm M, François B, Mercier E, Garbino J, Laterre P-F, Koch H, et al. 2014. Pseudomonas aeruginosa serotypes in nosocomial pneumonia: prevalence and clinical outcomes. Crit Care. 18(1):R17. doi:https://doi.org/10.1186/cc13697.
- Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, et al. 2012. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect. 18(3):268–281. doi:https://doi.org/10.1111/j.1469-0691.2011.03570.x.
- Mesaros N, Nordmann P, Plésiat P, Roussel-Delvallez M, Van Eldere J, Glupczynski Y, Van Laethem Y, Jacobs F, Lebecque P, Malfroot A, et al. 2007. Pseudomonas aeruginosa: resistance and therapeutic options at the turn of the new millennium. Clin Microbiol Infect. 13(6):560–578. doi:https://doi.org/10.1111/j.1469-0691.2007.01681.x.
- Milivojevic D, Sumonja N, Medic S, Pavic A, Moric I, Vasiljevic B, Senerovic L, Nikodinovic-Runic J. 2018. Biofilm-forming ability and infection potential of Pseudomonas aeruginosa strains isolated from animals and humans. Pathog Dis. 76(4):1–14. doi:https://doi.org/10.1093/femspd/fty041.
- Miyata S, Casey M, Frank DW, Ausubel FM, Drenkard E. 2003. Use of the Galleria mellonella caterpillar as a model host to study the role of the type III secretion system in Pseudomonas aeruginosa pathogenesis. Infect Immun. 71(5):2404–2413. doi:https://doi.org/10.1128/IAI.71.5.2404-2413.2003.
- Neidig A, Yeung AT, Rosay T, Tettmann B, Strempel N, Rueger M, Lesouhaitier O, Overhage J. 2013. TypA is involved in virulence, antimicrobial resistance and biofilm formation in Pseudomonas aeruginosa. BMC Microbiol. 13:77. doi:https://doi.org/10.1186/1471-2180-13-77.
- O’Toole GA, Kolter R. 1998. Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol. 30(2):295–304. doi:https://doi.org/10.1046/j.1365-2958.1998.01062.x.
- Pirnay JP, Bilocq F, Pot B, Cornelis P, Zizi M, Van Eldere J, Deschaght P, Vaneechoutte M, Jennes S, Pitt T, et al. 2009. Pseudomonas aeruginosa population structure revisited. PLoS One. 4:e7740. doi:https://doi.org/10.1371/journal.pone.0007740.
- Radó J, Kaszab E, Petrovics T, Pászti J, Kriszt B, Szoboszlay S. 2017. Characterization of environmental Pseudomonas aeruginosa using multilocus sequence typing scheme. J Med Microbiol. 66(10):1457–1466. doi:https://doi.org/10.1099/jmm.0.000589.
- Radhapriya P, Ramachandran A, Anandham R, Mahalingam S. 2015. Pseudomonas aeruginosa RRALC3 enhances the biomass, nutrient and carbon contents of Pongamia pinnata seedlings in degraded forest soil. PLoS One. 10:e0139881. doi:https://doi.org/10.1371/journal.pone.0139881.
- Roshani-Asl P, Rashidi N, Shokoohizadeh L, Zarei J. 2018. Relationship among antibiotic resistance, biofilm formation and lasB gene in Pseudomonas aeruginosa isolated from burn patients. Clin Lab. 64(9):1477–1484. doi: https://doi.org/10.7754/Clin.Lab.2018.180331.
- Rutherford V, Yom K, Ozer EA, Pura O, Hughes A, Murphy KR, Cudzilo L, Mitchel D, Hauser AR. 2018. Environmental reservoirs for exoS+ and exoU+ strains of Pseudomonas aeruginosa. Environ Microbiol Rep. 10(4):485–492. doi:https://doi.org/10.1111/1758-2229.12653.
- Salimi H, Yakchali B, Owlia P, Lari AR. 2010. Molecular epidemiology and drug susceptibility of Pseudomonas aeruginosa strains isolated from burn patients. Lab Med. 41:540–544. doi:https://doi.org/10.1309/LMNIJE31EDC1WAMP.
- Schmidt J, Müsken M, Becker T, Magnowska Z, Bertinetti D, Möller S, Zimmermann B, Herberg FW, Jänsch L, Häussler S. 2011. The Pseudomonas aeruginosa chemotaxis methyltransferase CheR1 impacts on bacterial surface sampling. Plos One. 6(3):e18184. doi:https://doi.org/10.1371/journal.pone.0018184.
- Schroeder M, Brooks BD, Brooks AE. 2017. The complex relationship between virulence and antibiotic resistance. Genes (Basel). 8(1): pii: E39. doi:https://doi.org/10.3390/genes8010039.
- Shaver CM, Hauser AR. 2004. Relative contributions of Pseudomonas aeruginosa ExoU, ExoS and ExoT to virulence in the lung. Infect Immun. 7(2):6969–6977. doi:https://doi.org/10.1128/IAI.72.12.6969-6977.2004.
- Shehata MMK, Sayed AA. 2011. Genetic diversity and twitching motility of Pseudomonas aeruginosa strains isolated from different origins. Arch Clin Microbiol. 2(5):4.
- Silby MW, Winstanley C, Godfrey SAC, Levy SB, Jackson RW. 2011. Pseudomonas genomes: diverse and adaptable. FEMS Microbiol Rev. 35(4):652–680. doi:https://doi.org/10.1111/j.1574-6976.2011.00269.x.
- Spilker T, Coenye T, Vandamme P, LiPuma JJ. 2004. PCR-based assay for differentiation of Pseudomonas aeruginosa from other Pseudomonas species recovered from cystic fibrosis patients. J Clin Microbiol. 42(5):2074–2079. doi:https://doi.org/10.1128/JCM.42.5.2074-2079.2004.
- Stepanović S, Vuković D, Dakić I, Savić B, Švabić-Vlahović M. 2000. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J Microbiol Methods. 40(2):175–179. doi:https://doi.org/10.1016/S0167-7012(00)00122-6.
- Strateva T, Mitov I. 2011. Contribution of an arsenal of virulence factors to pathogenesis of Pseudomonas aeruginosa infections. Ann Microbiol. 61(4):717–732. doi:https://doi.org/10.1007/s13213-011-0273-y.
- Taee SR, Khansarinejad B, Abtahi H, Najafimosleh M, Gzanavi-Rad E. 2014. Detection of algD, oprL and exoA genes by new specific primers as an efficient, rapid and accurate procedure for direct diagnosis of Pseudomonas aeruginosa strains in clinical samples. Jundishapur J Microbiol. 7(10):e 13583.
- Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH, Swaminathan B. 1995. Interpreting chromosomal DNA restriction patterns produced by pulsed- field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 33(9):2233–2239.
- Thrane SW, Taylor VL, Freschi L, Kukavica-Ibrulj I, Boyle B, Laroche J, Pirnay J-P, Lévesque RC, Lam JS, Jelsbak L. 2015. The widespread multidrug-resistant serotype O12 Pseudomonas aeruginosa clone emerged through concomitant horizontal transfer of serotype antigen and antibiotic resistance gene clusters. mBio. 6(5):e01396–15. doi:https://doi.org/10.1128/mBio.01396-15.
- Tokajian S, Timani R, Issa N, Araj G. 2012. Molecular characterization, multiple drug resistance and virulence determinants of Pseudomonas aeruginosa isolated from Lebanon. Br Microbiol Res J. 2:243–250. doi:https://doi.org/10.9734/BMRJ/2012/2217.
- Tsai CJ-Y, Loh JMS, Proft T. 2016. Galleria mellonella infection models for the study of bacterial diseases and for antimicrobial drug testing. Virulence. 7(3):214–229. doi:https://doi.org/10.1080/21505594.2015.1135289.
- Ullah W, Qasim M, Rahman H, Jie Y, Muhammad N. 2017. Beta-lactamase-producing Pseudomonas aeruginosa: phenotypic characteristics and molecular identification of virulence genes. J Chinese Med Assoc. 80(3):173–177. doi:https://doi.org/10.1016/j.jcma.2016.08.011.
- Van Duuren JBJH, Müsken M, Karge B, Tomasch J, Wittmann C, Häussler S, Brönstrup M. 2017. Use of single-frequency impedance spectroscopy to characterize the growth dynamics of biofilm formation in Pseudomonas aeruginosa. Sci Rep. 7. doi:https://doi.org/10.1038/s41598-017-05273-5.
- Velikova N, Kavanagh K, Wells JM. 2016. Evaluation of Galleria mellonella larvae for studying the virulence of Streptococcus suis. BMC Microbiol. 16:1–9. doi:https://doi.org/10.1186/s12866-016-0905-2.
- Yasmin S, Hafeez FY, Mirza MS, Rasul M, Arshad HMI, Zubair M, Iqbal M. 2017. Biocontrol of bacterial leaf blight of rice and profiling of secondary metabolites produced by rhizospheric Pseudomonas aeruginosa BRp3. Front Microbiol. 8. doi:https://doi.org/10.3389/fmicb.2017.01895.