Abstract
Aim: Chromones are promising for anticancer drug development. Methods & results: 12 chromone-based compounds were synthesized and tested against cancer cell lines. Compound 8 showed the highest cytotoxicity (LC50 3.2 μM) against colorectal cancer cells, surpassing 5-fluorouracil (LC50 4.2 μM). It suppressed colony formation, induced cell cycle arrest and triggered apoptotic cell death, confirmed by staining and apoptosis markers. Cell death was accompanied by enhanced reactive oxygen species formation and modulation of the autophagic machinery (autophagy marker light chain 3B (LC3B); adenosine monophosphate-activated protein kinase (AMPK); protein kinase B (PKB); UNC-51-like kinase (ULK)-1; and ULK2). Molecular docking and dynamic simulations revealed that compound 8 directly binds to ULK1. Conclusion: Compound 8 is a promising lead for autophagy-modulating anti-colon cancer drugs.
GRAPHICAL ABSTRACT
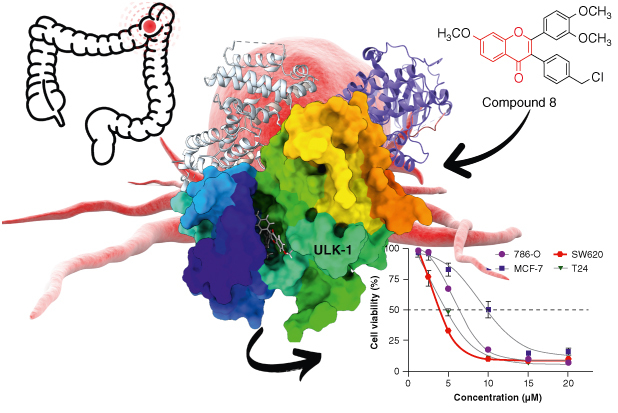
12 chromone-based compounds were synthesized and assessed for cytotoxicity against four human cancer cell lines i.e. kidney (786-0), breast (MCF-7), colorectal (SW620) and bladder (T-24) cancer cells.
Compound 8 exhibited the highest cytotoxic activity (LC50 3.2 μM) compared with the positive control 5-fluorouracil (5-FU, LC50 4.2 μM), especially against SW620 colorectal carcinoma cells.
Compound 8 suppressed colony formation, induced cell cycle arrests and triggered apoptotic cell death, confirmed by propidium iodide/annexin V staining and expression studies on apoptosis markers.
Enhanced formation of reactive oxygen species (ROS) and modulation of autophagic machinery (LC3B, AMPK, AKT, ULK1 and ULK2) were observed in response to compound 8.
A molecular docking and molecular dynamics (MD) simulations suggested a potential direct binding between compound 8 and ULK1.
Compound 8 represents a promising lead for a novel class of autophagy-modulating anti-colon cancer drugs.
Supplemental material
Supplemental data for this article can be accessed at https://doi.org/10.1080/17568919.2024.2363668
Acknowledgments
The authors thank the Universiti Kebangsaan Malaysia (UKM)-Malaysia for providing Discovery Studio®3.1 software and Chulalongkorn University-Thailand for MD simulation facilities and computing resources.
Author contributions
K Rullah contributed a key role in conceptualization and served as the project leader. NF Shamsudin and S-W Leong conducted the experiments and performed data analysis. A Koeberle and LK Wai contributed to designing the experiments, as well as preparing the manuscript. Additionally, MFFM Aluwi and SMM Faudzi carried out the docking simulations, while U Suriya and T Rungrotmongkol performed the MD simulations. The interpretation of data and preparation of the manuscript involved contributions from SL Chia, M Taher, MS Haris, HA Alshwyeh, AA Alosaimi, A Mediani, MA Ilowefah and D Islami.
Financial disclosure
This work was supported by the Ministry of Higher Education (MOHE) Malaysia through a Fundamental Research Grant Scheme (FRGS/1/2022/STG04/UIAM/02/2). The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Competing interests disclosure
The authors have no competing interests or relevant affiliations with any organization or entity with the subject matter or materials discussed in the manuscript. This includes employment, consultancies, honoraria, stock ownership or options, expert testimony, grants or patents received or pending, or royalties.
Writing disclosure
No writing assistance was utilized in the production of this manuscript.
