ABSTRACT
The objective of this study was to develop a reliable visual method for rapid detection of donkey components to protect consumers from commercial adulteration. Six specific loop-mediated isothermal amplification (LAMP) primers targeted at donkey mitochondrial cytochrome b gene were designed, the mitochondrial DNA extraction simplified, the LAMP reaction system optimized, the specificity verified with mitochondrial DNA of horse, pork, cow, sheep, chicken, duck, and rabbit as negative controls, the detection limit determined with gradient dilution of adulterated meat with donkey meat, and the visual LAMP method for detection of donkey-derived ingredient in common meat products established. The results showed that the modified mitochondrial DNA extraction method was simple and repeatable, and the visual LAMP method with 4-(2-pyridylazo)-resorcinol sodium salt as indicator can accurately and specifically detect the donkey meat in common meat products, 1% detection limit. The study provided a promising solution for facilitating the surveillance of the commercial adulteration in processed meat.
RESUMEN
El presente estudio se propuso desarrollar un método visual confiable que permitiera detectar rápidamente la presencia de componentes de carne de burro en carnes procesadas para proteger a los consumidores de la adulteración de alimentos comerciales. Con este fin se diseñaron seis cebadores LAMP específicos dirigidos al gen del citocromo B mitocondrial del burro, se simplificó la extracción de ADN mitocondrial, se optimizó el sistema de reacción LAMP y se verificó su especificidad con ADN mitocondrial de caballo, cerdo, vaca, oveja, pollo, pato y conejo que operaron como controles negativos. Además, el límite de detección se fijó mediante la dilución en gradiente de carne adulterada con carne de burro, estableciéndose el método visual LAMP para revelar la presencia de ingredientes derivados de carne de burro en productos cárnicos comunes. Los resultados obtenidos mostraron que el método de extracción de ADN mitocondrial modificado era simple y repetible, y que el método visual LAMP con sal sódica de 4-(2-piridilazo)-resorcinol como indicador sirve para detectar de manera precisa y específica la presencia de carne de burro en productos cárnicos comunes, con un límite de detección del 1%. El estudio proporcionó una solución prometedora para facilitar la vigilancia de la adulteración comercial de carnes procesadas.
1. Introduction
The food labels of meat and meat products are required to mark the meat source specifically to prevent adulteration in many countries, but the mixture of lower-price meat into high-price meat is still common, for example, adulterating pork or duck meat into beef or mutton or donkey meat, therefore, a reliable identification method of meat species is critical for surveillance of the commercial adulteration (Hong et al., Citation2017). The existing identification methods are mainly classified into protein-based approaches, for example, radioimmunoassay (Lowenstein et al., Citation2006), chromatography (Lozano et al., Citation2017; Pebriana et al., Citation2017; Wu et al., Citation2018) and Chemometrics-Assisted Shotgun Proteomics (Yuswan et al., Citation2018), and DNA-based approaches (Sheikha et al., Citation2017), for example, polymerase chain reaction (PCR) (Karabasanavar et al., Citation2017; Man et al., Citation2012; Mane et al., Citation2012), multi-PCR (Abuzinadah et al., Citation2015; Jia et al., Citation2016), real-time PCR (ÅAkalar & Kaynak, Citation2016; Herrero et al., Citation2013; Pegels et al., Citation2015; Sudjadi et al., Citation2016), PCR-RFLP (Bielikova et al., Citation2010), Biochip technology (Iwobi et al., Citation2011), forensically informative nucleotide sequencing (Rajpoot et al., Citation2017), and real-time PCR coupled melting curve analysis (Yuru et al., Citation2016). The structure of meat protein are usually destroyed by the processes such as shredding, cooking and roasting, therefore, the reliability of protein-based approaches is compromised, by comparison, the DNA-based approaches are more reliable, among which PCR is the most commonest assay, PCR, forensically informative nucleotide sequencing and melting curve for analyzing the adulterated meat product is very effective, but limited by the presence of PCR inhibitors in real biological samples and food samples, and meat products are the very complex matrix, mainly composed of proteins, lipids, pigments, enzymes, and other substances, which may interfere with PCR reactions (Wilson, Citation1997). It is still necessary to develop a rapid, sensitive, and cost-effective method for identification of adulterated meat.
Loop-mediated isothermal amplification (LAMP) has been developed to amplify nucleic acids specifically, sensitively, and rapidly under isothermal conditions (Notomi et al., Citation2000), the method has been developed to meat adulteration detection such as pork, duck, horse, cow, Jumbo Flying Squid, and so on (Deb et al., Citation2016; Li & Fan, Citation2016; Ran et al., Citation2015; Shi et al., Citation2017; Ye et al., Citation2017; Zahradnik et al., Citation2015), and the objective of this study is to develop a reliable visual LAMP method for rapid detection of donkey-derived ingredients to protect consumers from commercial adulteration.
2. Materials and methods
2.1. Design of LAMP primers
Targeting donkey mitochondrial cytochrome b gene (GenBank: FJ428360.1) (Yacoub & Sadek, Citation2017), a set of six LAMP primers were designed and selected with PrimerExplorer 5 and Oligo 7, the criteria of primer selection were ⊿G ≤ −4 Kcal/mol of F3/B3, FIP (F2 and F1 c)/BIP (B2 and B1 c), and LF/LB/, listed in and .
Table 1. LAMP primers for detecting donkey mitochondrial cytochrome B gene.
Tabla 1. Cebadores LAMP para detectar el gen del citocromo B mitocondrial de burro
Figure 1. Primer sequences and their location. Note: FIP (Forward inner primer) composed of F2 and F1 c; BIP (Backward inner primer) composed of B2 and B1 c.
Figura 1. Secuencias del cebador y su ubicación. Nota: FIP (Forward Inner Primer) compuesto de F2 y F1 c; BIP (Backward Inner Primer) compuesto de B2 y B1 c

2.2. Modification of mitochondrial DNA extraction
The meat of donkey, horse, pork, cow, sheep, chicken, duck, and rabbit were purchased from Xuchang Farm Product Market. The meat tissue or meat products of 200 mg was cut into small pieces, washed with 1 mL PBS buffer (PH7.9), dried with filter paper, grinded with 1 mL TE buffer (PH8.0) in ice bath, and centrifuged for 5 min at 1000 × g; the supernatant was transferred to a new tube and centrifuged for 5 min at 12000 × g, and the supernatant was discarded; the white precipitate was dissolved with 0.5 mL TE buffer (PH8.0), and the solution was centrifuged for 5 min at 1000 × g; the supernatant was transferred to a new tube and centrifuged for 5 min at 12000 × g, and the supernatant was discarded. The precipitate was dissolved with 50 μL TE buffer (PH8.0) and subjected to ultrasonication for 5 min, and the extract was directly used for LAMP reaction (Gargouri & Kacem, Citation2018).
2.3. Selection of LAMP reaction temperature
The LAMP was performed in a 20 μL reaction mixture containing 0.8 mM each of forward inner primer (FIP) and backward inner primer (BIP), 0.2 mM each of forward outer primer (F3) and backward outer primer (B3), 0.4 mM backward loop primer (LB), 0.4 mM forward loop primer (LF),1.0 mM dNTPs, 20 mM Tris-HCl (pH 8.8), 10 mM KCl, 10 mM (NH4)2SO4, 6 mM MgSO4, 0.5 mM MnSO4, 0.1% Triton X-100, 7.5% DMSO (Frackman et al., Citation1998), 0.5 mM 4-(2-pyridylazo)-resorcinol sodium salt (Shanghai Aladdin Biochemical Technology Co., Ltd. Shanghai, China), 1 pg donkey mitochondrial DNA template, and 8 U Bst 2.0 WarmStart DNA polymerase (New England Biolabs, Beverly, Mass., U.S.A.) (Wang, Citation2016). The reaction mixture was heated at 57°C, 58°C, 59°C, or 60°C for 60 min in water bath, and the reaction tubes were sunk into water. The LAMP products were subjected to electrophoresis on a 2.0% agarose gel, and visualized under UV light of the ChemiDoc MP Imaging System (Bio-Rad Laboratories, Inc. CA, USA) after ethidium bromide staining.
2.4. Sensitivity determination of the LAMP assay
The donkey was mixed with pork, according to proportion, the mass percent of donkey was 100%, 10%, 1%, and 0.1%, respectively, and their mitochondrial DNA were extracted according to above steps for sensitivity determination, and the mixture of donkey with horse, cow, sheep, chicken, duck, or rabbit was performed and their mitochondrial DNA was extracted in the same way. And the detection limit was determined with above LAMP reaction system at optimized temperature. In addition, fluorescent dyes calcein (Shanghai Aladdin Biochemical Technology Co., Ltd. Shanghai, China) had been used as the comparison with 4-(2-pyridylazo)-resorcinol sodium salt in the sensitivity determination, 0.5 mM MnSO4 and 0.5 mM 4-(2-pyridylazo)-resorcinol sodium salt were substituted by 6.25 mM calcein and 12.5 mM Mncl2 in above reaction system (Tomita et al., Citation2008).
2.5. Specificity determination of the LAMP assay
The mitochondrial DNA extracted from donkey, horse, pork, cow, sheep, chicken, duck, and rabbit were used for determining the specificity of our developed LAMP assay. In addition, the amount of mitochondrial DNA template used was 1 pg per reaction.
3. Results and analysis
3.1. Selection of the LAMP reaction temperature
The LAMP reaction with the newly designed primers was carried out at varying temperatures for 60 min, as shown in , resulted in amplification of all positive controls along with negative results for all negative control reactions. Given amplification efficiency, as indicated, 59°C was chosen as the most suitable reaction temperature.
Figure 2. LAMP reactions at varying temperatures. Note: A: 57°C; B: 58°C; C: 59°C; D: 60°C; PC: Positive control (donkey mitochondrial DNA template); NC: Negative control (mitochondrial DNA template substituted by TE buffer).
Figura 2. Reacciones de LAMP a diferentes temperaturas. Nota: A: 57°C; B: 58°C; C: 59°C; D: 60°C; PC: control positivo (plantilla de ADN mitocondrial de burro); NC: control negativo (plantilla de ADN mitocondrial sustituido por tampón TE)
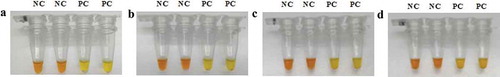

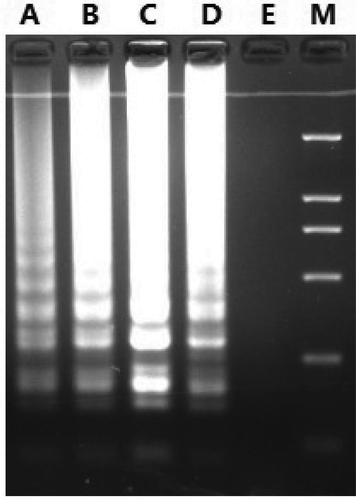
Figure 3. The optimal temperature of the LAMP assay for detection of donkey-derived ingredients. Note: A: 57°C; B: 58°C; C: 59°C; D: 60°C; E: Negative control (mitochondrial DNA template substituted by TE buffer); M: Marker DL2000.
Figura 3. Temperatura óptima del ensayo LAMP para la detección de ingredientes derivados de la carne de burro. Nota: A: 57°C; B: 58°C; C: 59°C; D: 60°C; E: control negativo (plantilla de ADN mitocondrial sustituido por tampón TE); M: marcador DL2000
3.2. Sensitivity of the LAMP assay
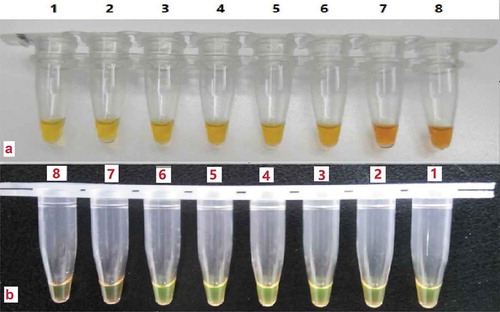
The sensitivity of the LAMP assay was determined with the mitochondrial DNA extracted from donkey meat mixed with different proportion of pork at 59°C for 60 min. The detection limit was found to be 1% donkey meat mixed with pork, as indicated. And the sensitivity of the LAMP assay for detection of donkey mixed with other species-derived meat determined to be 1% donkey meat, and the results of two indicator 4-(2-pyridylazo)-resorcinol sodium salt and calcein were consistent with.
Figure 4. Detection limit of the developed LAMP assay. Note: A: the LAMP assay with 4-(2-pyridylazo)-resorcinol sodium salt as indicator; B: the LAMP assay with calcein as indicator; 1, 2: 100% donkey meat; 3, 4: 10% donkey meat; 5, 6: 1% donkey meat; 7, 8: 0.1% donkey meat.
Figura 4. Límite de detección del ensayo LAMP desarrollado. Nota: A: ensayo LAMP con sal de sodio 4-(2-piridilazo)-resorcinol como indicador; B: ensayo LAMP con calceína como indicador; 1, 2: 100% carne de burro; 3, 4: 10% de carne de burro; 5, 6: 1% de carne de burro; 7, 8: 0.1% de carne de burro

3.3. Specificity of the LAMP assay
The specificity of the LAMP assay was tested with eight common animal-derived meat samples and as shown in ; both donkey meat samples were successfully detected, while meat derived from other species kept negative.
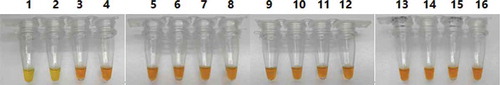
Figure 5. Specificity of the developed LAMP assay. Note: 1, 2: donkey; 3, 4: horse; 5, 6: pork; 7, 8: cow; 9, 10: sheep; 11, 12: chicken; 13, 14: duck; 15, 16: rabbit.
Figura 5. Especificidad del ensayo LAMP desarrollado. Nota: 1, 2: burro; 3, 4: caballo; 5, 6: carne de cerdo; 7, 8: vaca; 9, 10: oveja; 11, 12: pollo; 13, 14: pato; 15, 16: conejo
4. Discussion
Loop-mediated isothermal amplification (LAMP) was of high sensitivity (Notomi et al., Citation2000), so the aerosol pollution may form during the amplification, which leaded to false positive. In our experiment, the reaction tubes of LAMP were sunk in water, the aerosol that may leak was washed by water in water bath, and the false positive due to the aerosol pollution had been effectively reduced.
Most the existing mitochondrial DNA extraction kits extract the mitochondria from animal cells firstly, then extract and purify the DNA from the mitochondria, the operation is very complex, and the extraction time is more than 2 h. In the study, The animal mitochondrial DNA extraction were simplified via reduction of centrifugation steps, deletion of proteinase K lysis and addition of ultrasonication, and the animal DNA extraction time was shorten to less than 1 h. The purity of the mitochondrial DNA extracted with the simplified method was relatively low, but the results of the sensitivity determination indicated that the established visual LAMP assay can tolerate the inhibitors in the extracted mitochondrial DNA.
The chemical reaction indication dyes were generally classified as pH indicator dyes (Neutral Red, Phenol Red), calorimetric dyes (Malachite Green, Hydroxy Naphthol Blue, 4-(2-pyridylazo)-resorcinol Sodium Salt) and fluorescent dyes (Calcein, Ethidium Bromide, ROX Reference Dye, VeriPCR). The pH of LAMP system changed little with amplification reaction, the pH indicator dyes had poor performance; based on the LAMP reaction system and the chromogenic mechanism of different calorimetric dyes, 4-(2-pyridylazo)-resorcinol sodium salt was as indicator in the study; the fluorescent dye calcein had been used as a comparison (Tomita et al., Citation2008), and the results were consistent.
5. Conclusion
The animal mitochondrial DNA extraction was simplified. The extracted DNA purity was relatively low, however, the extraction efficiency of animal mitochondrial DNA was greatly improved, which also indicated that the LAMP assay was a robust DNA-based approach.
The visual LAMP method for detection of donkey-derived ingredients in common meat products was established in this study, and the detection limit was 1% donkey meat mixed into other species-derived meat. By comparison with other methods (ÅAkalar & Kaynak, Citation2016; Jia et al., Citation2016; Karabasanavar et al., Citation2017; Pebriana et al., Citation2017; Rajpoot et al., Citation2017; Sudjadi et al., Citation2016; Yuru et al., Citation2016), the developed LAMP assay had the advantages of cost effective, simplicity, high specificity, and sensitivity, which was suitable for surveillance of the commercial meat or meat products adulterated with donkey meat.
Disclosure statement
No potential conflict of interest was reported by the authors.
Additional information
Funding
References
- ÅAkalar, E., & Kaynak, A. (2016). Practical molecular detection method of beef and pork in meat and meat products by intercalating dye based duplex real-time polymerase chain reaction. International Journal of Food Properties, 19(1), 31–40. https://doi.org/doi.10.1080/10942912.2015.1017049
- Abuzinadah, O. H., Yacoub, H. A., El Ashmaoui, H. M., & Ramadan, H. A. (2015). Molecular detection of adulteration in chicken products based on mitochondrial 12S rRNA gene. Mitochondrial DNA, 26(3), 337–340. https://doi.org/10.3109/19401736.2013.840593
- Bielikova, M., Pangallo, D., & Turna, J. (2010). Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) as a molecular discrimination tool for raw and heat-treated game and domestic animal meats. Journal of Food and Nutrition Research, 49(3), 134–139. https://doi.org/doi.10.3168/jds.2009-2342
- Deb, R., Sengar, G. S., Singh, U., Kumar, S., Alyethodi, R. R., Alex, R., Raja, T. V., Das, A. K., & Prakash, B. (2016). Application of a loop-mediated isothermal amplification assay for rapid detection of cow components adulterated in buffalo milk/meat. Molecular Biotechnology, 58(12), 1–11. https://doi.org/10.1007/s12033-016-9984-4
- Frackman, S., Kobs, G., Simpson, D., & Storts, D. (1998). Betaine and DMSO: Enhancing agents for PCR. Promega Notes, 65, 27.
- Gargouri, H., & Kacem, H. H. (2018). Evaluation of alternative DNA extraction protocols for the species determination in turkey salami authentication tests. International Journal of Food Properties, 21(1), 733–745. https://doi.org/doi.10.1080/10942912.2017.1422263
- Herrero, B., Royo, L. J., Lago, F. C., Vieites, J. M., & Espiñeira, M. (2013). Authentication of male beef by multiplex fast real-time PCR. Food Additives & Contaminants: Part A, 30(2), 218–225. https://doi.org/doi.10.1080/19440049.2012.740164
- Hong, E., Lee, S. Y., Jeong, J. Y., Park, J. M., Kim, B. H., Kwon, K., & Chun, H. S. (2017). Modern analytical methods for the detection of food fraud and adulteration by food category. Journal of the Science of Food and Agriculture, 97(12), 3877–3896. https://doi.org/10.1002/jsfa.2017.97.issue-12
- Iwobi, A. N., Huber, I., Hauner, G., Miller, A., & Busch, U. (2011). Biochip technology for the detection of animal species in meat products. Food Anal Methods, 4(3), 389–398. https://doi.org/10.1007/s12161-010-9178-9
- Jia, X., Wei, Z., Mengru, Z., Yuanju, W., Tao, X., Xiaoqian, H., Yongfeng, Z., Suizhong, C., Lili, N., Hongping, Z., & Tao, Z. (2016). Molecular identification of adulteration in mutton based on mitochondrial 16S rRNA gene. Mitochondrial DNA Part A, 27(1), 628–632. https://doi.org/10.3109/19401736.2014.908377
- Karabasanavar, N., Girish, P. S., Kumar, D., & Singh, S. P. (2017). Detection of beef adulteration by mitochondrial D-loop based species-specific polymerase chain reaction. International Journal of Food Properties, 20(sup2), 2264–2271. https://doi.org/doi.10.1080/10942912.2017.1369103
- Li, Y. J., & Fan, J. Y. (2016). Rapid visual identification of bovine meat by loop mediated isothermal amplification combined with immunochromatographic strip. Biochip Journal, 11(1), 1–6. https://doi.org/doi.10.1007/s13206-016-1102-y
- Lowenstein, J. M., Reuther, J. D., Hood, D. G., Scheuenstuhl, G., Gerlach, S. C., & Ubelaker, D. H. (2006). Forensic Sci Int. Identification of Animal Species by Protein Radioimmunoassay of Bone Fragments and Bloodstained Stone Tools, 159(2–3), 182. https://doi.org/doi.10.1016/j.forsciint.2005.08.007
- Lozano, M., Rodríguez-Ulibarri, P., Echeverría, J. C., Beruete, M., Sorolla, M., & Beriain, M. J. (2017). Mid-infrared spectroscopy (MIR) for simultaneous determination of fat and protein content in meat of several animal species. Food Analytical Methods, 10(10), 1–9. https://doi.org/doi.10.1007/s12161-017-0879-1
- Man, Y. B. C., Mustafa, S., Mokhtar, N. F. K., Nordin, R., & Sazili, A. Q. (2012). Porcine-specific polymerase chain reaction assay based on mitochondrial D-loop gene for identification of pork in raw meat. International Journal of Food Properties, 15(1), 134–144. https://doi.org/doi.10.1080/10942911003754692
- Mane, B. G., Mendiratta, S. K., Tiwari, A. K., & Bhilegaokar, K. N. (2012). Detection of adulteration of meat and meat products with buffalo meat employing polymerase chain reaction assay. Food Anal Methods, 5(2), 296–300. https://doi.org/10.1007/s12161-011-9237-x
- Notomi, T., Okayama, H., Masubuchi, H., Yonekawa, T., Watanabe, K., Amino, N., & Hase, T. (2000). Loop-mediated isothermal amplification of DNA. Nucleic Acids Research, 28(12), e63. https://doi.org/doi.10.1097/RLU.0b013e3181f49ac7
- Pebriana, R. B., Rohman, A., Lukitaningsih, E., & Sudjadi. (2017). Development of FTIR spectroscopy in combination with chemometrics for analysis of rat meat in beef sausage employing three lipid extraction systems. International Journal of Food Properties, 20(sup2), 1995–2005. https://doi.org/doi.10.1080/10942912.2017.1361969
- Pegels, N., García, T., Martín, R., & González, I. (2015). Market analysis of food and feed products for detection of horse DNA by a Taqman real-time PCR. Food Anal Methods, 8(2), 489–498. https://doi.org/10.1007/s12161-014-9914-7
- Rajpoot, A., Kumar, V. P., Bahuguna, A., & Kumar, D. (2017). Forensically informative nucleotide sequencing (FINS) for the first time authentication of Indian Varanus species: Implication in wildlife forensics and conservation. Mitochondrial DNA Part A, 28(6), 892–900. https://doi.org/10.1080/24701394.2016.1202943
- Ran, G., Ren, L., Han, X., Liu, X., Li, Z., Pang, D., Ouyang, H., & Tang, X. (2015). Development of a rapid method for the visible detection of pork DNA in halal products by loop-mediated isothermal amplification. Food Anal Methods, 9(3), 1–6. https://doi.org/doi.10.1007/s12161-015-0246-z
- Sheikha, A. F. E., Mokhtar, N. F. K., Amie, C., Lamasudin, D. U., Isa, N. M., & Mustafa, S. (2017). Authentication technologies using DNA-based approaches for meats and halal meats determination. Food Biotechnology, 31(4), 281–315. https://doi.org/doi.10.1080/08905436.2017.1369886
- Shi, Y., Feng, Y., Xu, C., Xu, Z., Cheng, D., & Lu, Y. (2017). Loop-mediated isothermal amplification assays for the rapid identification of duck-derived ingredients in adulterated meat. Food Anal Methods, 10(7), 1–7. https://doi.org/10.1007/s12161-016-0767-0
- Sudjadi, Wardani, H. S., Sepminarti, T., & Rohman, A. (2016). Analysis of Porcine Gelatin DNA in a Commercial Capsule Shell Using Real-Time Polymerase Chain Reaction for Halal Authentication. International Journal of Food Properties. 19(9), 2127–2134. https://doi.org/doi.10.1080/10942912.2015.1110164
- Tomita, N., Mori, Y., Kanda, H., & Notomi, T. (2008). Loop-mediated isothermal amplification (lamp) of gene sequences and simple visual detection of products. Nature Protocols, 3(5), 877–882. https://doi.org/10.1038/nprot.2008.57
- Wang, D. (2016). Novel primers for increased specificity and sensitivity for the detection of by real-time LAMP. CyTA - Journal of Food, 14(1), 88–91. https://doi.org/doi.10.1080/19476337.2015.1048530
- Wilson, I. G. (1997). Inhibition and facilitation of nucleic acid amplification. Applied and Environmental Microbiology, 63(10), 3741–3751. https://doi.org/10.1089/oli.1.1997.7.523
- Wu, T., Zhong, N., & Yang, L. (2018). Identification of adulterated and non-adulterated Norwegian salmon using FTIR and an improved PLS-DA method. Food Anal Methods, 11(5), 1501–1509. https://doi.org/10.1007/s12161-017-1135-4
- Yacoub, H. A., & Sadek, M. A. (2017). Identification of fraud (with pig stuffs) in chicken-processed meat through information of mitochondrial cytochrome b. Mitochondrial DNA Part A, 28(6), 855–859. https://doi.org/10.1080/24701394.2016.1197220
- Ye, J., Feng, J., Dai, Z., Meng, L., Zhang, Y., & Jiang, X. (2017). Application of loop-mediated isothermal amplification (LAMP) for rapid detection of jumbo flying squid dosidicus gigas (D’Orbigny, 1835). Food Anal Methods, 10(5), 1452–1459. https://doi.org/10.1007/s12161-016-0700-6
- Yuru, T., Chao, J., Yuan, Y., Yan, J., Zhanhu, C., & Luqi, H. (2016). Molecular identification of antelope horn by melting curve analysis. Mitochondrial DNA Part A, 27(6), 3945–3951. https://doi.org/10.3109/19401736.2014.989500
- Yuswan, M. H., Aizat, W. M., Lokman, A. A., Desa, M. N. M., Mustafa, S., Junoh, N. M., Yuso, Z. N. B., Mohamed, R., Mohmad, Z., & Lamasudin, D. U. (2018). Chemometrics-assisted shotgun proteomics for establishment of potential peptide markers of non-halal pork (Sus scrofa) among halal beef and chicken. Food Anal Methods, 11(12), 3505–3515. https://doi.org/doi.10.1007/s12161-018-1327-6
- Zahradnik, C., Martzy, R., Mach, R. L., Krska, R., Farnleitner, A. H., & Brunner, K. (2015). Loop-mediated isothermal amplification (lamp) for the detection of horse meat in meat and processed meat products. Food Anal Methods, 8(6), 1576–1581. https://doi.org/10.1007/s12161-014-0072-8
