Abstract
The plastid genome of Lysidice brevicalyx was successfully assembled using Illumina sequencing reads for the first time. The complete plastid genome of L. brevicalyx is a circular structure of 159,084 bp with a GC content of 36.4%. It comprises a large single-copy (LSC) region of 87,783 bp, a small single-copy (SSC) region of 19,557 bp, and two inverted repeat regions (IRA and IRB) of 25,872 bp, each. The plastome of L. brevicalyx contains a total of 128 genes, including 83 protein-coding genes, 37 tRNAs, and 8 rRNAs. The phylogenetic analysis strongly supports the monophyly of Lysidice. This study provides the first complete plastid genome sequence of L. brevicalyx and contributes to our understanding of the molecular characteristics and evolutionary relationships of this plant species.
Introduction
The genus Lysidice (Fabaceae) consists of two species, viz. Lysidice rhodostegia Hance 1867 from China and Vietnam and Lysidice brevicalyx C. F. Wei 1983 endemic to southern China (Hou Citation2010). L. brevicalyx is a valuable tree species used for building materials due to its white-yellow wood () and has been used as an alternative medicine for the treatment of fractures and traumatic bleeding in China (Gao et al. Citation2007). Additionally, the roots, stems, and leaves of L. brevicalyx have been used for medicinal purposes by local communities (Wu et al. Citation2010, Yang et al. Citation2019). In this study, we report the first assembly and annotation of the complete plastid genome of L. brevicalyx () using second-generation sequencing technology.
Figure 1. Plant image of Lysidice brevicalyx. The image showing flowers are actinomorphic, with purple flag and wing petals, an evenly pinnate leaf, flattened, twisted pods. This photo was taken by youpai zeng.

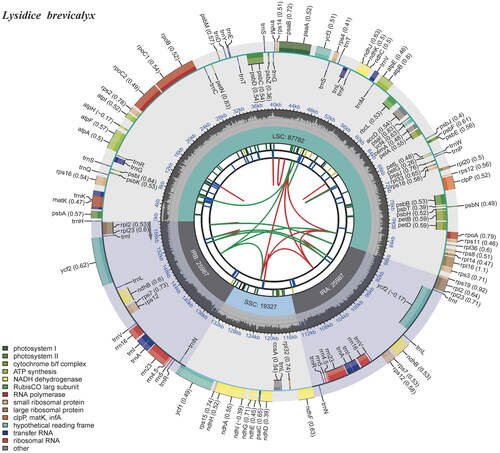
Figure 2. Gene map of the plastid genome of Lysidice brevicalyx. From the center outward, the first track indicates the dispersed repeats; the second track shows the long tandem repeats as short blue bars; the third track shows the short tandem repeats or microsatellite sequences as short bars with different colors; the fourth track shows small single-copy (SSC), inverted repeat (IRA and IRB), and large single-copy (LSC) regions. The GC content along the genome is plotted on the fifth track; the genes are shown on the sixth track.
Materials and methods
Fresh leaves of L. brevicalyx were collected from the South China Botanical Garden in Guangzhou City, Guangdong Province, China (N 23°11'12.62′', E 113°21'51.15′'). A voucher specimen (Q. Lai LaiQ036) was deposited in IBSC (contact person: Qiang Lai, [email protected]). Total genomic DNA was extracted from the fresh leaves using a combination of the improved cetyltrimethylammonium bromide (CTAB) method (Doyle and Doyle Citation1987, Yang et al. Citation2014) and the Dneasy Plant Mini Kit extraction. Quality monitoring was performed using a Qubit 3.0 fluorometer. Genomic DNA was sheared to prepare a PCR-free library with an insert size of 150 bp. High-throughput sequencing was conducted using the Illumina Hiseq X-Ten system, generating a total of 2.3 GB of pair-end reads. The plastid genome was assembled using the GetOrganelle v1.7.7.0 (Jin et al. Citation2020), and the unique genes of the L. brevicalyx plastid genome were annotated using CPGAVAS2 web service (Shi et al. Citation2019). A gene graphical map of the plastid genome was constructed using CPGVIEW (http://www.1kmpg.cn/ cpgview) (Liu et al. Citation2023). Then final plastid genome of L. brevicalyx was submitted to NCBI Gene Bank with an accession number of OQ808806. Firstly, a total of 19 plastid genomes were downloaded from NCBI Gene Bank, 74 protein-coding genes shared by all genomes were screened. Subsequently, MAFFT v7.313 (Katoh et al. Citation2019) was used for separate alignment of each gene, and maximum likelihood phylogenies were inferred using IQ-TREE (Minh et al. Citation2022) under the GTR + F + I + G4 model for 5000 ultrafast bootstraps.
Results and discussion
The newly assembled plastid genome of Lysidice brevicalyx is 159,084 bp in length. The sequencing coverage depths of the genome ranged from 7× to 1873× with a mean coverage depth of 1195.94×, indicating the reliability of the genome assembly. The plastid gene structure of L. brevicalyx is a circular molecule (), consisting of four parts: a large single-copy region (LSC) of 87,783 bp, a small single-copy region (SSC) of 19,557 bp, and two inverted repeat regions (IRA and IRB), each 25,872 bp. The plastome contains a total of 128 genes, including 83 protein-coding genes, 37 tRNAs, 8 rRNAs. Among the protein-coding genes, 14 protein-coding genes (atpF, ndhA, ndhB, petB, petD, rpl16, rpoC1, rps16, trnA-UGC, trnG-UCC, trnI-GAU, trnK-UUU, trnL-UAA, and trnV-UAC) contain one intron and three genes (clpP, rps12, and ycf3) have two introns. Three small-exon genes (petB, petD, and rps16) and one trans-spliced gene (rps12) were verified to be corrected and annotated with multiple sequence alignment.
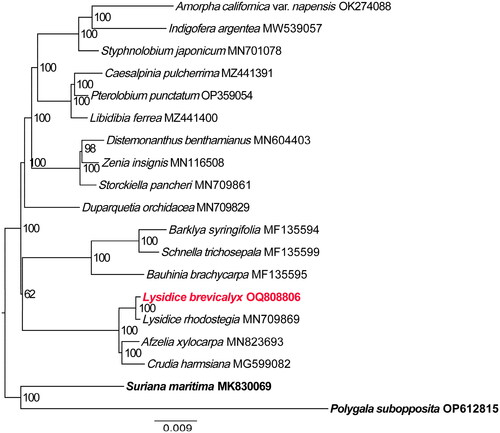
For the construction of the Maximum Likelihood (ML) phylogenetic tree, a total of 19 complete plastid genomes of fabales were used (). The result showed that L. brevicalyx is closely related to L. rhodostegia, Afzelia xylocarpa (Kurz) Craib, and Crudia harmsiana De Wil, and all of them formed a clade with high supported value (BP = 100%). These findings are consistent with previous research (Zhao et al. Citation2021). The clade comprising subfamilies Cercidoideae and Detarioideae is sister to the remaining legumes, and Duparquetioideae and Dialioideae are successive sisters to the clade of Papilionoideae and Caesalpinioideae.
Figure 3. The phylogenetic position for Lysidice brevicalyx according to the ML phylogenetic tree constructed based on74 plastid genomes. The following sequences were used: Polygala subopposita OP612815 and suriana maritima MK830069 (Lai et al. Citation2019b) as outgroup, Crudia harmsiana MG599082 (Tosso et al. Citation2018), Afzelia xylocarpa MN823693 (Zhang et al. Citation2020a), Lysidice rhodostegia MN709869 (Zhang et al. Citation2020b), bauhinia brachycarpa MF135595 (Wang et al. Citation2018), schnella trichosepala MF135599 (Wang et al. Citation2018) Barklya syringifolia MF135594 (Wang et al. Citation2018), duparquetia orchidacea MN709829 (Zhang et al. Citation2020b), storckiella pancheri MN709861 (Zhang et al. Citation2020b), zenia insignis MN116508 (Lai et al. Citation2019a) distemonanthus benthamianus MN604403 (Demenou et al. Citation2020), libidibia ferrea MZ441400 (Aecyo et al. Citation2021), pterolobium punctatum OP359054 (Zhang et al. Citation2020b), caesalpinia pulcherrima MZ441391 (Aecyo et al. Citation2021) styphnolobium japonicum MN701078 (Shi and Liu Citation2019), indigofera argentea MW539057 amorpha californica var. napensis OK274088 (Agudelo et al. Citation2022) The sequences used for the tree structure are coding sequences, and the bootstrap support values are shown on the nodes. The scale bar represents the numbers of substitutions per site.
Conclusion
This study presents the first report of the plastid genome of Lysidice brevicalyx, and adds to the limited number of plastid genomes reported for the genus Lysidice. The phylogenetic analysis revealed that Lysidice forms an independent sister clade to other species, providing novel insights into the phylogenetic relationships within the Leguminosae family. The genetic resource information generated in this study will be valuable for further investigations into the biology and evolutionary history of Lysidice.
Ethical approval
The material involved in the article does not involve ethical conflicts. Field studies complied with local legislation, and appropriate permissions were granted before the samples were collected from the South China Botanical Garden.
Authors’ contributions
In this research, Jianxin Li conceived and designed the study. She has completed the data analysis and the first draft of the paper. Qiang Lai collected the samples. Ying Meng, Ze-Long Nie and Tie-Yao Tu have been responsible for supervising the experimental design, data analysis, and the writing and revision of the manuscript. All authors approved the final version and agreed to be accountable for all aspects of the work.
Supplemental Material
Download MS Word (4.9 MB)Disclosure statement
No potential conflict of interest was reported by the authors.
Data availability statement
The complete plastid genome sequence of L. brevicalyx has been deposited in the NCBI GenBank database under the accession number OQ808806 (these numbers were automatically generated by NCBI and refer to the same sample). The associated BioProject, SRA, and Bio-Sample numbers are PRJNA955444, SRR24167046, and SAMN34178680 respectively.
Additional information
Funding
References
- Aecyo P, Marques A, Huettel B, Silva A, Esposito T, Ribeiro E, Leal IR, Gagnon E, Souza G, Pedrosa-Harand A, et al. 2021. Plastome evolution in the Caesalpinia group (Leguminosae) and its application in phylogenomics and populations genetics. Planta J. 254(2):27. doi:10.1007/s00425-021-03655-8.
- Agudelo ID, Aldaco G, Brito-Pizano A, Chavez KG, Cortina KG, Flores J, Fuentes A, Garcia AN, Garcia A, Gonzalez-Martinez D, et al. 2022. The complete chloroplast genome of the threatened Napa False Indigo Amorpha californica var. napensis Jeps. 1925 (Fabaceae) from Northern California, USA. Mitochondrial DNA B Resour. 7(1):283–285. doi:10.1080/23802359.2022.2029605.
- Demenou BB, Migliore J, Heuertz M, Monthe FK, Ojeda DI, Wieringa JJ, Dauby G, Albreht L, Boom A, Hardy OJ, et al. 2020. Plastome phylogeography in two African rain forest legume trees reveals that Dahomey Gap populations originate from the Cameroon volcanic line. Mol Phylogenet Evol. 150:106854. doi:10.1016/j.ympev.2020.106854.
- Doyle JJ, Doyle JL. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull. 19:11–15.
- Gao S, Liu J, Fu G-M, Hu Y-C, Yu S-S, Fan L-H, Yu D-Q, Qu J. 2007. Resveratrol/phloroglucinol glycosides from the roots of Lysidice rhodostegia. Planta Med. 73(2):163–166. doi:10.1055/s-2006-951770.
- Hou CDZDD. 2010. Flora of China [M]. Beijing: Science Press.
- Jin J-J, Yu W-B, Yang J-B, Song Y, dePamphilis CW, Yi T-S, Li D-Z. 2020. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21(1):241. doi:10.1186/s13059-020-02154-5.
- Katoh K, Rozewicki J, Yamada KD. 2019. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform. 20(4):1160–1166. doi:10.1093/bib/bbx108.
- Lai Q, Tu T, Zhang D. 2019a. The complete plastid genome of Zenia insignis Chun (Leguminosae). Mitochondrial DNA B Resour. 4(2):2926–2927. doi:10.1080/23802359.2019.1660266.
- Lai Q, Zhu C, Gu S, Tu T, Zhang D. 2019b. Complete plastid genome of Suriana maritima L. (Surianaceae) and its implications in phylogenetic reconstruction of Fabales. J Genet. 98:109. doi:10.1007/s12041-019-1157-3.
- Liu S, Ni Y, Li J, Zhang X, Yang H, Chen H, Liu C. 2023. CPGView: a package for visualizing detailed chloroplast genome structures. Mol Ecol Resour. 23(3):694–704. doi:10.1111/1755-0998.13729.
- Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Von Haeseler A, Lanfear R. 2022. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 39(5):1530–1534. doi:10.1093/molbev/msaa015.
- Shi L, Chen H, Jiang M, Wang L, Wu X, Huang L, Liu C. 2019. CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 47(W1):W65–W73. doi:10.1093/nar/gkz345.
- Shi Y, Liu B. 2019. Complete chloroplast genome sequence of Sophora japonica ‘JinhuaiJ2’ (Papilionaceae), an important traditional chinese herb. Mitochondrial DNA B Resour. 5(1):319–320. doi:10.1080/23802359.2019.1703588.
- Tosso F, Hardy OJ, Doucet J-L, Daïnou K, Kaymak E, Migliore J. 2018. Evolution in the Amphi-Atlantic tropical genus Guibourtia (Fabaceae, Detarioideae), combining NGS phylogeny and morphology. Mol Phylogenet Evol. 120:83–93. doi:10.1016/j.ympev.2017.11.026.
- Wang Y-H, Wicke S, Wang H, Jin J-J, Chen S-Y, Zhang S-D, Li D-Z, Yi T-S. 2018. Plastid genome evolution in the early-diverging Legume subfamily Cercidoideae (Fabaceae). Front Plant Sci. 9:138. doi:10.3389/fpls.2018.00138.
- Wu X-F, Hu Y-C, Yu S-S, Jiang N, Ma J, Tan R-X, Li Y, Lv H-N, Liu J, Ma S-G, et al. 2010. Lysidicins F − H, Three New Phloroglucinols from Lysidice rhodostegia. Org Lett. 12(10):2390–2393. doi:10.1021/ol100735f.
- Yang JB, Li DZ, Li HT. 2014. Highly effective sequencing whole chloroplast genomes of angiosperms by nine novel universal primer pairs. Mol Ecol Resour. 14(5):1024–1031. doi:10.1111/1755-0998.12251.
- Yang L-M, Yin H, Zhang M-J, Zhang M, Huang X-F. 2019. Three new phenolic glycosides from the roots of Lysidice rhodostegia. Phytochem Lett. 33:125–128., doi:10.1016/j.phytol.2019.08.002.
- Zhang J, Li Y, Heng S, Seab K, Wang Y. 2020a. The complete chloroplast genome sequence of Afzelia xylocarpa. Mitochondrial DNA B Resour. 5(1):762–763. doi:10.1080/23802359.2020.1715857.
- Zhang R, Wang Y-H, Jin J-J, Stull GW, Bruneau A, Cardoso D, De Queiroz LP, Moore MJ, Zhang S-D, Chen S-Y, et al. 2020b. Exploration of plastid phylogenomic conflict yields new insights into the deep relationships of Leguminosae. Syst Biol. 69(4):613–622. doi:10.1093/sysbio/syaa013.
- Zhao Y, Zhang R, Jiang K-W, Qi J, Hu Y, Guo J, Zhu R, Zhang T, Egan AN, Yi T-S, et al. 2021. Nuclear phylotranscriptomics and phylogenomics support numerous polyploidization events and hypotheses for the evolution of rhizobial nitrogen-fixing symbiosis in Fabaceae. Mol Plant. 14(5):748–773. doi:10.1016/j.molp.2021.02.006.
