Abstract
Objectives
MicroRNAs (miRNAs) play essential roles in the development of COPD. In this study, we aimed to identify and validate potential miRNA biomarkers in frequent and non-frequent exacerbators of COPD patients using bioinformatic analysis.
Materials and methods
The candidate miRNA biomarkers in COPD were screened from Gene Expression Omnibus (GEO) dataset and identified using GEO2R online tool. Then, we performed bioinformatic analyses including target prediction, gene ontology (GO), pathway enrichment analysis and construction of protein–protein interaction (PPI) network. Furthermore, the expression of the identified miRNAs in peripheral blood monocular cells (PBMCs) of COPD patients was validated using quantitative real-time polymerase chain reaction (qRT-PCR).
Results
MiR-23a, miR-25, miR-145 and miR-224 were identified to be significantly downregulated in COPD patients compared with healthy controls. GO analysis showed the four miRNAs involved in apoptotic, cell differentiation, cell proliferation and innate immune response. Pathway analysis showed that the targets of these miRNAs were associated with p53, TGF-β, Wnt, VEGF and MAPK signal pathway. In healthy controls, the miR-25 and miR-224 levels were significantly decreased in smokers compared with nonsmokers (P<0.001 and P<0.05, respectively). In COPD patients, the levels of miR-23a, miR-25, miR-145 and miR-224 were associated with Global Initiative for Chronic Obstructive Lung Disease (GOLD) stages. Notably, miR-23a and miR-145 were significantly elevated in non-frequent exacerbators compared with frequent exacerbators (P<0.05), and miR-23a showed higher area under the receiver–operator characteristic curve (AUROC) than miR-145 (0.707 vs 0.665, P<0.05).
Conclusion
MiR-23a, miR-25, miR-145 and miR-224 were associated with the development of COPD, and miR-23a might be a potential biomarker for discriminating the frequent exacerbators from non-frequent exacerbators.
Introduction
COPD is characterized by persistent airflow limitation due to airway and/or alveolar abnormalities that are usually caused by exogenous exposure to noxious particles or gases.Citation1 Cigarette smoking is the major risk factor for the development of COPD. In 2010, COPD is the fourth leading cause of death worldwide, but it is predicted to be the third by 2020.Citation2 Acute exacerbation of COPD (AECOPD), which is generally characterized by the aggravated dyspnea and the increased volumes of phlegm and phlegm purulence, had an in-hospital mortality of 4.3%–7.7%.Citation3–Citation5 However, there are still no strong and effective strategies to predict the occurrence of AECOPD. Therefore, biomarkers for predicting the development and prognosis of COPD are urgently required.
MicroRNAs (miRNAs) are endogenous, approximately 20–25 nt, noncoding RNAs that can target mRNAs for the cleavage of translational repression.Citation6 miRNAs have been demonstrated to participate in the pathogenesis and development of COPD. Fujita et alCitation7 reported that miR-210 contributed to abnormal airway remodeling by suppressing autophagy and promoting myofibroblast differentiation. Osei et alCitation8 demonstrated that miR-146a-5p involved in the epithelial–fibroblast cross talk and the blockage of miR-146a-5p might induce the chronic inflammation in COPD-derived primary human lung fibroblasts. Furthermore, miRNAs have also been demonstrated to be promising and effective biomarkers for the diagnosis of COPD. In 2012, Akbas et alCitation9 first reported the downregulation of serum miR-20, miR-28-3p, miR-34c-5p and miR-100 and the upregulation of miR-7 in COPD patients compared with healthy controls, and they suggested that these miRNAs might provide potential biomarkers for therapeutic strategy. Another group reported that the ratio of serum miR-20a to miR-181a was associated with the early stage of COPD in asymptomatic heavy smokers.Citation10 In addition, Wang et alCitation11 also demonstrated that the levels of miR-145-5p, miR-338-3p and miR-3620-3p were related to the severity of COPD. However, these preliminary results were usually limited by small sample size and have not been confirmed using large-scale dataset.
With the development of high-throughput microarray and sequencing technology, several public resources have been established, and the Gene Expression Omnibus (GEO) of National Center for Biotechnology Information (NCBI) is the largest one. Bioinformatic analyses based on the GEO database provide valuable information for searching biomarkers in various diseases.Citation12–Citation16 However, little data have been reported on the bioinformatics-based identification of potential miRNA biomarkers for COPD patients. Therefore, we aimed to identify and validate potential miRNA biomarkers in frequent and non-frequent exacerbators of COPD patients using bioinformatic analysis.
Materials and methods
Selection of candidate miRNAs and target prediction
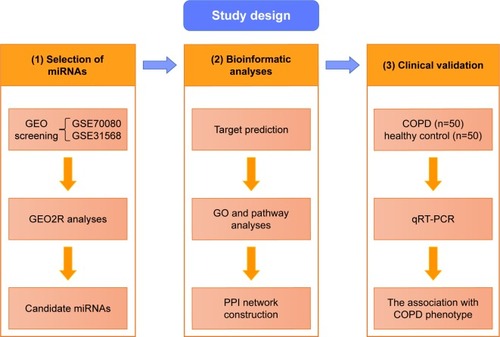
The general overview of the study design is shown in . Shared blood miRNA profiles of COPD patients were searched in GEO datasets.Citation17 GSE70080 and GSE31568 were selected, and the dysregulated miRNAs of COPD patients and healthy controls were intersected. GEO2RCitation18 was performed to compare the original submitter-supplied processed data tables using the GEO query and limma R packages from the bioconductor project. Generally, GEO2R is a well-designed tool for avoiding the false positivity by handling the various types of data and performing multiple testing corrections on P-values.
Figure 1 The general overview of the study design.
Abbreviations: GEO, Gene Expression Omnibus; GO, gene ontology; miRNAs, microRNAs; PBMCs, peripheral blood monocular cells; PPI, protein–protein interaction; qRT-PCR, quantitative real-time polymerase chain reaction.
The accuracy for the predicted targets of candidate miRNAs was improved by the following software: PicTarCitation19, miRDBCitation20 and TargetScan.Citation21 Validated targets were acquired from TarBase.Citation22 Briefly, we took the intersection of the three series of predicted targets and validated the targets using TarBase.
Gene ontology (GO) and pathway enrichment analysis of target genes
GeneCodisCitation23 is a web-based tool for comprehensive analysis of gene annotations, encompassing GO, Panther and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, InterPro Motifs and Transcription Factors, etc. GO annotates genes and gene products including molecular function (MF), cellular component (CC) and biological process (BP).Citation24 Panther and KEGG are bioinformatic resources for genomic and gene functional information.Citation25,Citation26 The significant enrichment results were accepted at a threshold of ≥2 gene counts with a P-value <0.05.
Construction of protein–protein interaction (PPI) network
Venn diagram was carried out for finding potential targets of the candidate miRNAs (Venny 2.1).Citation27 The interactions of the potential target genes were predicted by the Search Tool for the Retrieval of Interacting Genes (STRING) database.Citation28 Cytoscape (version 3.5.1; http://cytoscape.org/) is an open source software which is capable of integrating the high-throughput data and other types of molecular data into a unitized conceptual framework.Citation29 The values of gene interactions predicted by STRING were imported into Cytoscape to visualize the resulting PPIs and to identify hub genes among potential targets.
Patients and healthy controls
A total of consecutive 50 COPD patients were included during October 2016 to February 2017 in Qilu Hospital of Shandong University, as well as 50 well age- and gender-matched healthy subjects were recognized as controls. The inclusive criteria for COPD patients were set as forced expiratory volume in 1 s/forced vital capacity (FEV1/FVC)<0.7 according to the Global Initiative for Chronic Obstructive Lung Disease (GOLD) guideline. The classification of the severity stage for COPD was the following: GOLD I, forced expiratory volume in the first second of expiration for predicted values (FEV1%pre) ≥80%; GOLD II, 50%≤FEV1%pre<80%; GOLD III, 30%≤FEV1%pre<50%; GOLD IV, FEV1%pre<30%.Citation1 Frequent exacerbator was identified as COPD patient with two or more episodes of acute exacerbations during preceding 1 year, while non-frequent exacerbator as COPD patient with <2 episodes of acute exacerbations during preceding 1 year. Meanwhile, all patients did not undergo acute exacerbation during the nearly past 3 months. Exclusion criteria included the existences of other chronic lung diseases, tumors, unstable cardiovascular diseases, diabetes, nervous system diseases and liver and kidney diseases. The study was approved by the medical ethics committee of Qilu Hospital of Shandong University, and the written informed consent form was obtained from each patient.
Peripheral blood mononuclear cell (PBMC) separation and RNA extraction
The fasting peripheral vein blood (4 mL) was collected in EDTA tubes, and PBMCs were obtained through the Ficoll (Solarbio Life Sciences, Beijing, People’s Republic of China) isolation method. Total RNA containing small RNA was extracted from PBMCs by using the Trizol reagent (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol. RNA purity and concentration were evaluated by the ratio of absorbance at OD260/OD280 through ultraviolet spectrophotometer.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Quantitative real-time polymerase chain reaction (qRT-PCR) was performed with Bulge-loop™ miRNA qRT-PCR Starter kit on RT-PCR system (Thermo Fisher Scientific) according to the following protocol: reverse transcription PCR: 42°C 60 min, 70°C 10 min, 4°C ∞; qPCR: 95°C for 10 min, followed by 40 cycles of 95°C for 2 s, 60°C for 20 s and then 70°C for 10 s. Melt curve analysis was performed; the nuclear acid melting temperatures were 70°C–95°C; the heating rate was 0.5°C/time; holding time was 5 s/time; then, the Ct was recorded. The primers of these miRNAs and U6 were obtained from RiboBio Corporation (Guangzhou, People’s Republic of China). The fold change of each miRNA expression was calculated using the ΔΔCT method.
Statistical analyses
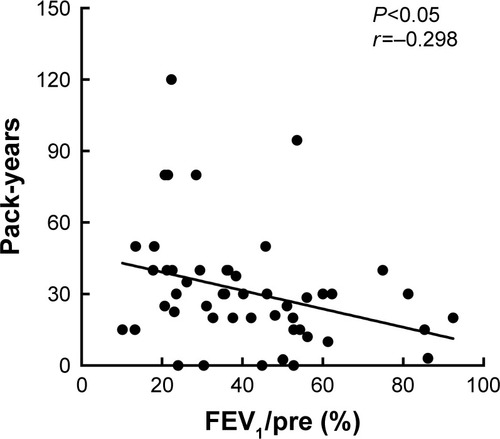
The demography and clinical data are presented as mean ± SD. The difference of gender composition between healthy controls and COPD patients was analyzed by Pearson’s chi-square test, and the differences for continuous variables were analyzed by Mann–Whitney U-test. The relationship between FEV1%pre and pack-years of COPD patients was analyzed by Pearson’s correlation analysis. The Mann–Whitney U-test was employed to compare the expression levels of miRNAs between the two groups. Receiver–operator characteristic (ROC) curve analysis and the area under the ROC curve (AUROC) were used to evaluate the specificity and sensitivity of miRNAs, with the larger AUROC value showing the higher diagnostic values. All statistical analyses were performed using the SPSS 20.0 software (IBM Corporation, Armonk, NY, USA) and the GraphPad Prism 6 (Graph-Pad Software, Inc., La Jolla, CA, USA). Differences with P-value of <0.05 were considered statistically significant.
Results
Identification of candidate miRNAs and prediction of targets
In GSE31568 and GSE70080, COPD samples and healthy controls were entered into GEO2R analysis to identify the candidate miRNAs. MiR-23a, miR-25, miR-145 and miR-224 have been selected as candidate miRNAs (P<0.05). The target genes were predicted from both prediction algorithms (TargetScan, miRDB and PicTar) and experimentally supported targets from TarBase. Eventually, we gained a total of 3114 target genes of the candidate miRNAs.
GO, KEGG and Panther pathway functional analyses
GO enrichment analysis demonstrated that the main annotations for the MF of the 3114 target genes included DNA-dependent regulation of transcription, signal transduction, multicellular organismal development, apoptotic process, transmembrane transport, cell differentiation, cell proliferation, innate immune response, platelet activation, response to stress, response to hypoxia, cell adhesion, metabolic process, cell migration, lipid metabolic process, oxidation–reduction process, inflammatory response, cytokine-mediated signaling pathway, etc ().
Figure 2 Bioinformatic analysis of predicted target genes of candidate miRNAs.
Abbreviations: BP, biological process; GO, gene ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; miRNAs, microRNAs.
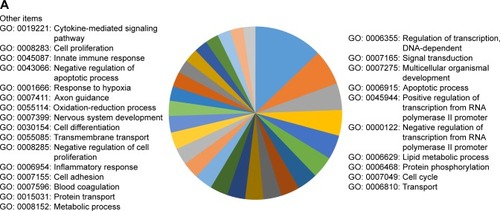
KEGG and Panther pathway analysis of these target genes presented p53 signal pathway, TGF-β signal pathway, Wnt signal pathway, VEGF signal pathway, MAPK signal pathway, oxidative stress response, Notch signal pathway, etc, indicating their potential roles for the pathogenesis of COPD (). The most enriched items of KGEE and Panther pathways according to P-values are presented in .
Table 1 KEGG and Panther pathway analysis of target genes
PPI network construction of potential targets
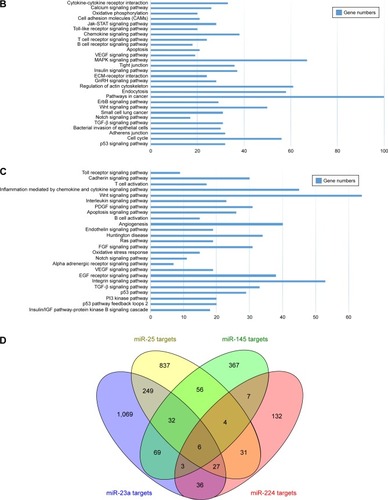
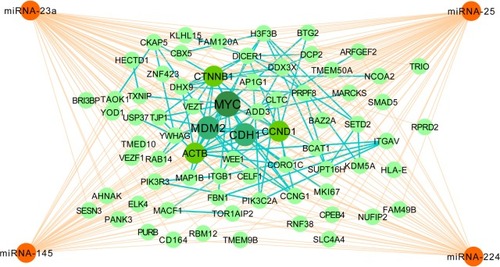
The Venn diagram of 3114 predicted genes is shown in . There are 32 common target genes (FBN1, ADD3, AHNAK, AP1G1, BAZ2A, etc) for miR-23a, miR-25 and miR-145, four genes (BTG2, TMEM9B, ELK4, ACTB) for miR-25, miR-145 and miR-224, 27 genes (MARCKS, YWHAG, SLC4A4, ARFGEF2, BIR3BP, etc) for miR-23, miR-25, miR-224, three genes (ZNF423, KDM5A, USP37) for miR-23, miR-145 and miR-224, as well as six genes (CCNG1, CCND1, CELF1, DDX3X, MAP1B and NUFIP1) for all the four miRNAs. Furthermore, the PPI network of the 72 identified target genes is shown in . In , ACTB, MYC, CTNNB1, CDH1 and CCND1 are shown as hub target genes according to the counts of interacting protein.
Figure 3 Interaction of miRNAs and targets and interaction of protein–protein network.
Abbreviation: miRNAs, microRNAs.

Basic demographic characteristics of the COPD patients and healthy controls
In this current study, the validation set included a total of 50 COPD patients (four for GOLD I, 14 for GOLD II, 15 for GOLD III and 17 for GOLD IV) and 50 healthy controls (32 nonsmokers and 18 smokers). The demographic and clinical characteristics of COPD patients and healthy controls are presented in and . The post-bronchodilator (BD) FEV1%pre of each GOLD grade (mean ± SD) is shown as follows: GOLD I, 86.29%±4.63%; GOLD II, 56.47%±6.51%; GOLD III, 38.74%±5.44%; GOLD IV, 19.83%±7.03%. In COPD patients, there were 18 non-frequent exacerbators and 32 frequent exacerbators. FEV1%pre was demonstrated to be negatively correlated with pack-years in COPD patients (P<0.05, r=−0.298; ). Between healthy controls and COPD patients, there were no significant differences in age and body mass index (BMI; all P>0.05) as well as significant difference in smoking status (P<0.001).
Table 2 The demographic and clinical characteristics of clinical investigation subjects
Table 3 The clinical characteristics of COPD patients with different GOLD stages
Validation of the identified miRNAs in PBMCs of healthy and COPD subjects
The levels of miR-23a, miR-25, miR-145 and miR-224 in PBMCs were found to be significantly lower in COPD patients than those in healthy controls (). Interestingly, miR-23a and miR-145 were downregulated in frequent exacerbators compared with non-frequent exacerbators (P<0.05). In healthy controls, the levels of miR-25 and miR-224 were upregulated in nonsmokers compared with those in smokers (P<0.001, P<0.05, respectively).
Figure 5 qRT-PCR validation of four candidate miRNAs in PBMCs of participants.
Abbreviations: GOLD, Global Initiative for Chronic Obstructive Lung Disease; miRNAs, microRNAs; PBMCs, peripheral blood monocular cells; qRT-PCR, quantitative real-time polymerase chain reaction.
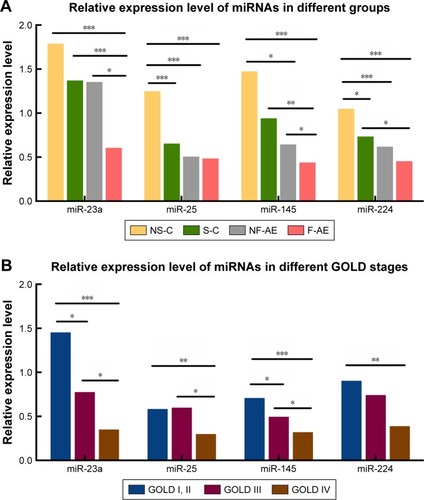
As shown in , the levels of miR-23a and miR-145 in GOLD I and GOLD II were significantly higher than GOLD III (P<0.05) and GOLD IV (P<0.001), as well as the same trend of miR-23a and miR-145 in GOLD III compared with GOLD IV (P<0.05). There was no significant difference in miR-25 between GOLD I and GOLD II compared with GOLD III (P>0.05), while there was a significantly higher expression of miR-25 in GOLD I and GOLD II and GOLD III compared with GOLD IV (P<0.01 and P<0.05, respectively). For the level of miR-224, there was no significant difference in GOLD I and GOLD II compared with GOLD III (P>0.05), as well as the same trend between GOLD III and GOLD IV (P>0.05). Of note, we demonstrated the significant higher expression of miR-224 in GOLD I than GOLD IV (P<0.01).
ROC curve analyses of miR-23a and miR-145 as biomarkers in non-frequent and frequent COPD exacerbators
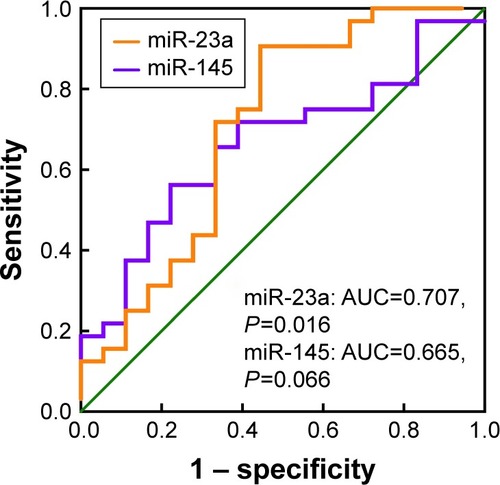
ROC analysis was performed to estimate the diagnostic accuracy of miR-23a and miR-145 as biomarkers for identifying frequent and non-frequent exacerbators (). The AUROC was 0.707 (95% CI: 0.543–0.880) with a sensitivity of 71.88% and specificity of 66.67% for miR-23a (P=0.016), as well as 0.665 (95% CI: 0.5123–0.8175) with a sensitivity of 65.63% and specificity of 66.67% for miR-145 (P=0.066).
Figure 6 ROC curve of miRNA-23a and miRNA-145 in COPD patients.
Abbreviations: ROC, receiver–operator characteristic; miRNAs, microRNAs; AUC, area under the curve.
Discussion
AECOPD is an acute event of respiratory symptoms with serious impact on quality of life of COPD patients.Citation30 However, little data have been reported on exploring the miRNA biomarkers for identifying frequent and non-frequent exacerbators of COPD patients. In the current study, we first identified the profile of miR-23a, miR-25, miR-145 and miR-224 and provided the functional pathways for those miRNAs using bioinformatic methods. Furthermore, we demonstrated that miR-23a and miR-145 were significantly different in non-frequent and frequent exacerbators using validated cohort. Finally, we suggested that miR-23a might be the potential and promising biomarker for discriminating frequent exacerbators from non-frequent exacerbators of COPD patients.
In recent years, increasing evidences showed that miRNAs contributed to the development and progression of COPD patients. MiR-199a-5p was demonstrated to be a critical regulator for unfolded protein response in alpha-1 antitrypsin-deficient monocytes.Citation31 Blockage of miR-503 has been reported to augment the release of vascular endothelial growth factor from lung fibroblasts of COPD patients.Citation32 MiR-218-5p, which was significantly downregulated in COPD patients compared with nonsmokers, was recognized as a protective factor for COPD.Citation33 Furthermore, Shen et alCitation34 reported that miR-483-5p was capable of abrogating the TGF-β-mediated cell proliferation in pulmonary epithelial and fibroblast cell lines. In addition, Shen et alCitation35 demonstrated that miR-149-3p might attenuate the inflammatory response via TLR-4/NF-κB signaling pathway in COPD patients.
In the current study, we first identified that the profiles of miR-23a, miR-25, miR-145 and miR-224 were down-regulated in COPD patients than healthy controls, indicating their potential roles in the development of COPD. Previously, miR-23a, miR-25, miR-145 and miR-224 have been found to be abnormally expressed in idiopathic pulmonary hypertension,Citation36 asthmaCitation37,Citation38 and lung cancer.Citation39–Citation41 MiR-23a was reported to be associated with the loss of muscle force during the AECOPD,Citation42 and the mechanism might be that miR-23a could regulate TLR2/MyD88/NF-κB pathway by targeting TLRs.Citation43 MiR-25 was demonstrated to mediate the phenotype of human airway smooth muscle cells (ASMCs) when exposing to IL-1β, TNF-α and IFN-γ.Citation44 MiR-145 can modulate the Th1/Th2 balance in asthma by regulating Runx3.Citation37 In the study by Hu et al,Citation40 miR-145 inhibited the TGF-β-induced epithelial–mesenchymal transition (EMT) by negatively regulating SMAD3 expression in human non-small cell lung cancer (NSCLC) cells. MiR-224 was reported to inhibit cell proliferation and metastasis in NSCLC A549 cell lines.Citation41 Shen et alCitation45 reported that miR-224 could downregulate gene expression of ACADM and ALDH2, inhibit the formation of triglyceride and influence lipid metabolism and apoptosis. Therefore, it is hypothesized that the profiles of miR-23a, miR-25, miR-145 and miR-224 might be associated with the inflammation, oxidative stress, immune imbalance, EMT, cell proliferation, apoptosis and lipid metabolism of COPD.
We have performed the GO and pathway analysis to predict the possible biological function and signal pathways of the target genes in the pathogenesis of COPD. Our results demonstrated that miR-23a, miR-25, miR-145 and miR-224 may play important roles in inflammation, oxidative stress, lipid metabolic disorders, cell proliferation and apoptosis. Enriched pathways included p53 signal pathway, TGF-β signal pathway, Wnt signal pathway, VEGF signal pathway, MAPK signal pathway, oxidative stress and Notch signal pathway. Tilley et alCitation46 reported the lower expression of NOTCH3 gene in smokers than nonsmokers, suggesting its potential interaction with smoke. Besides, TGF-β signal pathway involved in the development of lung, inflammation and airway remodeling of COPD. Another study showed that smoking-mediated airway inflammation could lead to oxidative stress and airway remodeling by activating MMP-9 and TGF-β/Smads pathway.Citation47 Therefore, we believe that our identified candidate miRNAs might contribute to the inflammation, airway mucus hypersecretion, oxidative stress and airway remodeling via those enriched pathways. However, the hypothesis based on the bioinformatic analyses is urgently needed to be confirmed in the future study.
In the current study, GSE70080 and GSE31568 were selected to perform the differential miRNA expression profile of COPD patients and healthy controls. It is worth noting that the two GEO Series also contain the gene information of lung cancer.Citation48,Citation49 Using the same GSE datasets, Halvorsen et alCitation48 revealed six circulating miRNAs (miR-429, miR-205, miR-200b, miR-203, miR-125b and miR-34b) for the early diagnosis of NSCLC patients, and Keller et alCitation49 reported some potential miRNA biomarkers for distinguishing lung cancer from COPD. Our current study mainly compared the data from COPD patients and healthy controls and demonstrated that miR-23a, miR-25, miR-145 and miR-224 were associated with the development of COPD.
Frequent exacerbation phenotype is one of the main phenotypes in COPD patients,Citation50,Citation51 and biomarkers for identifying the frequent and non-frequent exacerbators are urgently needed. In the previous study, urinary desmosine was reported to be a useful biomarker for identifying frequent exacerbators of COPD patients.Citation52 Besides, the upregulation of IL-1 pathway mediators (IRAK2, IRAK3, PELI1 and IL1R1)Citation53 was reported to be relevant to frequent exacerbations of COPD. Fu et alCitation54 demonstrated that serum IL-6 and sputum IL-1β were associated with the frequent exacerbations. However, miRNA biomarkers for identifying the frequent exacerbators of COPD have not been reported to date. Using the clinical samples, we demonstrated that miR-23a and miR-145 were significantly expressed between non-frequent and frequent exacerbators of COPD. Furthermore, ROC curve showed that miR-23a might be the potential and promising biomarker for discriminating frequent exacerbators from non-frequent exacerbators.
However, there are some limitations in our study. First, we performed bioinformatic analysis using two different groups and platforms, which might generate variances. Usually, there are two commonsense methods to avoid the variances: the first method is for performing quality control within each group using the techniques of normalization, principal component analysis (PCA) and cluster, then merging all the groups with raw data after quality control and eventually performing differential expression profiling in the merged group.Citation55–Citation58 However, the quality control might be quite disordered especially for the dataset with relatively small samples and inevitably generate poor repeatability. The second method is for first performing differential expression profiling in each group, respectively, and then choosing the common intersection. The P-value would be assigned for all miRNAs in the ranked lists to re-rank these miRNAs and decide their significance.Citation59–Citation64 This method might be more feasible for the dataset with relatively small samples or platform heterogeneity. Therefore, considering the real spirit of our current study, we have selected the second method to avoid the variances. Second, we still need to investigate the functional mechanism of the target genes and pathways in the future study, and the prognostic value of miR-23a should be confirmed in the prospective, multicenter, large-scale cohort.
Conclusion
We first identified the profile of miR-23a, miR-25, miR-145 and miR-224 and provided the functional pathways for those miRNAs using bioinformatic methods. Furthermore, we demonstrated that miR-23a and miR-145 were significantly different in non-frequent and frequent exacerbators using validated cohort. Finally, we suggested that miR-23a might be the potential and promising biomarker for discriminating frequent exacerbators from non-frequent exacerbators of COPD patients. However, the functional mechanism of the target genes and the prognostic value of miR-23a should be confirmed in the future study.
Acknowledgments
This study was supported by the grants from the Science and Technology Foundation of Shandong Province (grant number: 2007GG3002008, 2015GGH318002 and 2015GSF118064). Xiao Liu and Jingge Qu are the co-first authors.
Disclosure
The authors report no conflicts of interest in this work.
References
- Global Strategy for the Diagnosis, Management and Prevention of COPD, Global Initiative for Chronic Obstructive Lung Diseases (GOLD) 2016 Available from: http://goldcopd.org/Accessed Octobor 20, 2017
- LozanoRNaghaviMForemanKGlobal and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010Lancet201238098592095212823245604
- PereraPNArmstrongEPSherrillDLSkrepnekGHAcute exacerbations of COPD in the United States: inpatient burden and predictors of costs and mortalityCOPD20129213114122409371
- JohannesdottirSAChristiansenCFJohansenMBHospitalization with acute exacerbation of chronic obstructive pulmonary disease and associated health resource utilization: a population-based Danish cohort studyJ Med Econ201316789790623621504
- de Miguel-DiezJJimenez-GarciaRHernandez-BarreraVTrends in hospital admissions for acute exacerbation of COPD in Spain from 2006 to 2010Respir Med2013107571772323421969
- BartelDPMicroRNAs: genomics, biogenesis, mechanism, and functionCell2004116228129714744438
- FujitaYArayaJItoSSuppression of autophagy by extracellular vesicles promotes myofibroblast differentiation in COPD pathogenesisJ Extracell Vesicles201542838826563733
- OseiETFlorez-SampedroLTasenaHmiR-146a-5p plays an essential role in the aberrant epithelial-fibroblast cross-talk in COPDEur Respir J2017495160253828546273
- AkbasFCoskunpinarEAynaciEOltuluYMYildizPAnalysis of serum micro-RNAs as potential biomarker in chronic obstructive pulmonary diseaseExp Lung Res201238628629422686440
- XieLWuMLinHAn increased ratio of serum miR-21 to miR-181a levels is associated with the early pathogenic process of chronic obstructive pulmonary disease in asymptomatic heavy smokersMol Biosyst20141051072108124556821
- WangMHuangYLiangZPlasma miRNAs might be promising biomarkers of chronic obstructive pulmonary diseaseClin Respir J201610110411125102970
- WangYXueDLiYThe long noncoding RNA MALAT-1 is a novel biomarker in various cancers: a meta-analysis based on the GEO database and literatureJ Cancer201678991100127313790
- HuangYZhuJLiWSerum microRNA panel excavated by machine learning as a potential biomarker for the detection of gastric cancerOncol Rep20183931338134629286167
- HuTXiJIdentification of COX5B as a novel biomarker in high-grade glioma patientsOnco Targets Ther2017105463547029180880
- TianXZhuXYanTDifferentially expressed lncRNAs in gastric cancer patients: a potential biomarker for gastric cancer prognosisJ Cancer20178132575258628900495
- ManchiaMPirasISHuentelmanMJPattern of gene expression in different stages of schizophrenia: down-regulation of NPTX2 gene revealed by a meta-analysis of microarray datasetsEur Neuropsychopharmacol201727101054106328732597
- BarrettTSuzekTOTroupDBNCBI GEO: mining millions of expression profiles–database and toolsNucleic Acids Res200533 Database issueD562D52615608262
- BarrettTWilhiteSELedouxPNCBI GEO: archive for functional genomics data sets–updateNucleic Acids Res201341 Database issueD991D99523193258
- KrekAGrünDPoyMNCombinatorial microRNA target predictionsNat Genet200537549550015806104
- WangXImproving microRNA target prediction by modeling with unambiguously identified microRNA-target pairs from CLIP-ligation studiesBioinformatics20163291316132226743510
- AgarwalVBellGWNamJWBartelDPPredicting effective microRNA target sites in mammalian mRNAsElife20154e05005
- VlachosISParaskevopoulouMDKaragkouniDDIANA-TarBase v7.0: indexing more than half a million experimentally supported miRNA:mRNA interactionsNucleic Acids Res201543 Database issueD153D15925416803
- Nogales-CadenasRCarmona-SaezPVazquezMGeneCodis: interpreting gene lists through enrichment analysis and integration of diverse biological informationNucleic Acids Res200937 Web Server issueW317W32219465387
- Gene Ontology ConsortiumThe gene ontology (GO) project in 2006Nucleic Acids Res200634 Database issueD322D32616381878
- MiHDongQMuruganujanAGaudetPLewisSThomasPDPANTHER version 7: improved phylogenetic trees, orthologs and collaboration with the Gene Ontology ConsortiumNucleic Acids Res201038 Database issueD204D21020015972
- KanehisaMGotoSKEGG: Kyoto Encyclopedia of Genes and GenomesNucleic Acids Res2000281273010592173
- OliverosJC2007–2015Venny. An interactive tool for comparing lists with Venn’s diagrams Available from: http://bioinfogp.cnb.csic.es/tools/venny/index.html
- JensenLJKuhnMStarkMSTRING 8 – a global view on proteins and their functional interactions in 630 organismsNucl Acids Res200937 Database issueD412D41618940858
- ShannonPMarkielAOzierOCytoscape: a software environment for integrated models of biomolecular interaction networksGenome Res200313112498250414597658
- WedzichaJASeemungalTACOPD exacerbations: defining their cause and preventionLancet2007370958978679617765528
- HassanTCarrollTPBuckleyPGmiR-199a-5p silencing regulates the unfolded protein response in chronic obstructive pulmonary disease and alpha1-antitrypsin deficiencyAm J Respir Crit Care Med2014189326327324299514
- IkariJNelsonAJObaidJReduced microRNA-503 expression augments lung fibroblast VEGF production in chronic obstructive pulmonary diseasePLoS One2017129e018403928880936
- ConickxGMestdaghPAvila CobosFMicroRNA profiling reveals a role for microRNA-218-5p in the pathogenesis of chronic obstructive pulmonary diseaseAm J Respir Crit Care Med20171951435627409149
- ShenZTangWGuoJSunSmiR-483-5p plays a protective role in chronic obstructive pulmonary diseaseInt J Mol Med201740119320028534971
- ShenWLiuJZhaoGRepression of Toll-like receptor-4 by microRNA-149-3p is associated with smoking-related COPDInt J Chron Obstruct Pulmon Dis20171270571528260877
- SarrionIMilianLJuanGRole of circulating miRNAs as biomarkers in idiopathic pulmonary arterial hypertension: possible relevance of miR-23aOxid Med Cell Longev2015201579284625815108
- QiuYYZhangYWQianXFBianTmiR-371, miR-138, miR-544, miR-145, and miR-214 could modulate Th1/Th2 balance in asthma through the combinatorial regulation of Runx3Am J Trans Res20179731843199
- SuojalehtoHLindstromIMajuriMLAltered microRNA expression of nasal mucosa in long-term asthma and allergic rhinitisInt Arch Allergy Immunol2014163316817824513959
- HsuYLHungJYChangWAHypoxic lung cancer-secreted exosomal miR-23a increased angiogenesis and vascular permeability by targeting prolyl hydroxylase and tight junction protein ZO-1Oncogene201736344929494228436951
- HuHXuZLiCMiR-145 and miR-203 represses TGF-beta-induced epithelial-mesenchymal transition and invasion by inhibiting SMAD3 in non-small cell lung cancer cellsLung Cancer201697879427237033
- ZhuDChenHYangXDecreased microRNA-224 and its clinical significance in non-small cell lung cancer patientsDiagn Pathol2014919825410592
- DuanYZhouMXiaoJPrediction of key genes and miRNAs responsible for loss of muscle force in patients during an acute exacerbation of chronic obstructive pulmonary diseaseInt J Mol Med20163851450146228025995
- GuXGaoYMuDGFuEQMiR-23a-5p modulates mycobacterial survival and autophagy during mycobacterium tuberculosis infection through TLR2/MyD88/NF-kappaB pathway by targeting TLR2Exp Cell Res20173542717728327409
- KuhnARSchlauchKLaoRHalaykoAJGerthofferWTSingerCAMicroRNA expression in human airway smooth muscle cells: role of miR-25 in regulation of airway smooth muscle phenotypeAm J Respir Cell Mol Biol201042450651319541842
- ShenBPanQYangYmiR-224 affects mammary epithelial cell apoptosis and triglyceride production by downregulating ACADM and ALDH2 genesDNA Cell Biol2017361263327918674
- TilleyAEHarveyBGHeguyADown-regulation of the notch pathway in human airway epithelium in association with smoking and chronic obstructive pulmonary diseaseAm J Respir Crit Care Med2009179645746619106307
- ChurgACosioMWrightJLMechanisms of cigarette smoke-induced COPD: insights from animal modelsAm J Physiol Lung Cell Mol Physiol20082944L612L63118223159
- HalvorsenARBjaanaesMLeBlancMA unique set of 6 circulating microRNAs for early detection of non-small cell lung cancerOncotarget2016724372503725927191990
- KellerALeidingerPBauerAToward the blood-borne miRNome of human diseasesNat Methods201181084184321892151
- HurstJRVestboJAnzuetoASusceptibility to exacerbation in chronic obstructive pulmonary diseaseN Engl J Med2010363121128113820843247
- TashkinDPFrequent exacerbations of chronic obstructive pulmonary disease – a distinct phenotype?N Engl J Med2010363121183118420843256
- KimCKoYKimSHUrinary desmosine is associated with emphysema severity and frequent exacerbation in patients with COPDRespirology201823217618128905464
- BainesKJFuJJMcDonaldVMGibsonPGAirway gene expression of IL-1 pathway mediators predicts exacerbation risk in obstructive airway diseaseInt J Chron Obstruct Pulmon Dis20171254155028223794
- FuJJMcDonaldVMBainesKJGibsonPGAirway IL-1beta and Systemic Inflammation as Predictors of Future Exacerbation Risk in Asthma and COPDChest2015148361862925950204
- ShangguanHTanSYZhangJRBioinformatics analysis of gene expression profiles in hepatocellular carcinomaEur Rev Med Pharmacol Sci201519112054206126125269
- KongXDLiuNXuXJBioinformatics analysis of biomarkers and transcriptional factor motifs in Down syndromeBraz J Med Biol Res2014471083484125118625
- GuoYBaoYMaMYangWIdentification of key candidate genes and pathways in colorectal cancer by integrated bioinformatical analysisInt J Mol Sci2017184E72228350360
- ParkCYuNChoiIKimWLeeSlncRNAtor: a comprehensive resource for functional investigation of long non-coding RNAsBioinformatics201430172480248524813212
- SongESongWRenMIdentification of potential crucial genes associated with carcinogenesis of clear cell renal cell carcinomaJ Cell Biochem Epub20171211
- SongYJLiGHeJHGuoYYangLBioinformatics-based identification of microRNA-regulated and rheumatoid arthritis-associated genesPLoS One2015109e013755126359667
- ZhengLQWangRChiSMLiCXMatrix metalloproteinase 1: a better biomarker for squamous cell carcinoma by multiple microarray analysesG Ital Dermatol Venereol Epub20171215
- LiHWangHSunZGuoQShiHJiaYThe clinical and prognostic value of polo-like kinase 1 in lung squamous cell carcinoma patients: immunohistochemical analysisBiosci Rep Epub2017719
- LinZFShenXYLuFZRuanZHuangHLZhenJReveals new lung adenocarcinoma cancer genes based on gene expressionEur Rev Med Pharmacol Sci20121691249125623047510
- YangYLiHHouSHuBLiuJWangJDifferences in gene expression profiles and carcinogenesis pathways involved in cisplatin resistance of four types of cancerOncol Rep201330259661423733047