Abstract
Background
LARGE1 plays a pivotal role in glycosylation of alpha-Dystroglycan (α-DG) and is aberrantly downregulated in cell lines originating from epithelium-derived cancers including lung cancer. However, the expression of LARGE1 and its clinical significance in NSCLC are not clear.
Materials and Methods
The data were collected from the TCGA database to investigate LARGE1 expression in stage I–III NSCLC and explore its associations with clinicopathological parameters and overall survival of patients. The prognostic role of LARGE1 was examined in subgroups according to clinical features and treatments. The results were validated in external cohorts from the NCBI GEO database. Gene Set Enrichment Analysis (GSEA) was performed to investigate the potential molecular mechanisms during LARGE1 alteration in NSCLC.
Results
LARGE1 was aberrantly downregulated in NSCLC compared with adjacent tissues and normal lung tissues and in tumors with advanced stage compared with early stage. There was only a trend of association between high LARGE1 with OS in multivariate analysis. Surprisingly, high LARGE1 was significantly associated with improved OS in a subgroup of the patients with adjuvant chemotherapy (ACT) and a significant interaction between LARGE1 expression and ACT was found. Improved OS after ACT was also found in patients with high LARGE1 compared to those with low LARGE1. When combining LARGE1 expression and ACT, compared with patients with non-ACT, HR of low LARGE1/ACT was 0.592 (95% CI=0.432–0.813, P=0.0012), and HR of high LARGE1/ACT was 0.124 (95% CI=0.031–0.505, P=0.0036). The results were verified in two external cohorts from the GEO database. GSEA indicated that LARGE1 might downregulate cell cycle pathway to improve ACT sensitivity and subsequently the prognosis in NSCLC.
Conclusion
High LARGE1 can be used to identify the patients with resected stage I–III NSCLC most likely to benefit from adjuvant chemotherapy.
Introduction
Lung cancer is one of the most common malignant tumors and the leading cause of cancer-related deaths worldwide.Citation1 In 2020, it is expected that there will be 228,820 cases of lung cancer and 135,720 related deaths in the United States.Citation1 There are two main histological types of lung cancer: small cell lung cancer and non-small cell lung cancer (NSCLC).Citation2 NSCLC accounts for approximately 85% of lung cancer cases and can further be divided into lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC).Citation2 The 5-year survival rate of all patients with lung cancer is only about 19%.Citation1,Citation3
Surgical resection is the main and most effective choice for NSCLC patients, but only 20% of patients are suitable for surgery due to late diagnosisCitation4 and postoperative tumor recurrence is the major cause of treatment failure.Citation5–Citation7 Platinum-based adjuvant chemotherapy is one of the main means to improve the overall survival (OS) of patients with NSCLC after surgical resection, especially for patients with advanced tumorsCitation8 after several previous randomized clinical trials demonstrated ACT is associated with statistically significant survival advantages in patients with completely resected NSCLC.Citation9–Citation13 However, a meta-analysis of these studies suggested that ACT only provides an absolute benefit of 5.4% in 5-year OSCitation14 due to primary and acquired drug resistance.Citation15 And the remaining 85%–95% of patients do not get benefit from ACT but suffer the ACT toxicity. Thus, it is critical to identify NSCLC patients who will not benefit from ACT.
LARGE1, encoding a bifunctional glycosyltransferase, participates in glycosylation of α-DG by synthesizing and transferring the laminin-binding repeating units of 3GlcA-1-3Xyl-1 onto α-DG.Citation16 LARGE1 downregulation has been found to underlie abnormal glycosylation of α-DG in epithelium-derived cancer cells lines derived from breast, cervical, and lung cancers and exogenous expression of LARGE1 restores normal glycosylation and laminin binding of α-DG, leading to enhanced cell adhesion and reduced cell migration in vitro.Citation17 Exogenous expression of LARGE1 has also been found to restore α-DG matriglycan and laminin binding in rhabdomyosarcoma.Citation18
Currently, the clinical significance of LARGE1 in lung cancer is not clear. In the present study, we utilized the large dataset of NSCLC patients from TCGA database to investigate the expression of LARGE1 and its clinical significance and further validated the results in external cohorts. In addition, we preliminarily explored the potential molecular mechanisms related to LARGE1 in NSCLC carcinogenesis.
Materials and Methods
Expression of LARGE1 mRNA Expression in NSCLC
TCGA TARGET GTEx dataset, which contains gene expression data in tumor tissues, adjacent tissues, and normal lung tissues, was downloaded from Xenabrowser.netCitation19 and used to investigate LARGE1 mRNA expression in NSCLC tumor tissues, adjacent tissues, and normal lung tissues.
Kaplan–Meier Plotter
Kaplan-Meier plotter (KM plotter),Citation20 a web-based tool, is capable of assessing the effect of 54k genes (mRNA, miRNA, protein) on survival in 21 cancer types with the data from the databases including GEO, EGA, and TCGA. This tool consisted of 2,437 lung cancer cases with gene profile (mRNA) data determined by microarray. Of the 2,437 lung cases, 1,927 cases had OS information. We used the tool to initially analyze the association of LARGE1 expression with OS in lung cancer with Auto select best cutoff value and the JetSet best probe set (215543_s_at) for LARGE1.
Analysis of NSCLC Data from TCGA
To further investigate the associations of LARGE1 mRNA levels with clinicopathological features and OS in NSCLC, RNA-Seq data and corresponding clinical data of the NSCLC patients were downloaded from TCGA database by Xenabrowser.net.Citation19 The correlations between LARGE1 expression and baseline clinical characteristics in NSCLC patients were examined by the Chi-square test or Fisher’s exact test as appropriate. Kaplan-Meier plot curves were used to explore the association of LARGE1 expression (optimal cutoff value) with OS in overall NSCLC patients. Univariate/multivariate Cox proportional hazards regression models were also used to explore the effects of LARGE1 mRNA levels on OS in overall and subgroups stratified by age, gender, smoking history, tumor stage, margin status, and adjuvant chemotherapy (ACT) treatment. Furthermore, LARGE1 expression and ACT were also combined to classify NSCLC patients to observe their OS difference in Kaplan-Meier curves and Cox proportional hazards regression models.
Validation
To validate the results from the TCGA cohort, external cohort studies were searched in NCBI GEO database. The datasets in which: 1) the subjects were resected NSCLC patients; (2) the patients were treated with adjuvant chemotherapy and without chemotherapy; and 3) contained follow-up information (OS time and status) and contained LARGE1 expression levels analyzed by microarray or high throughput sequencing were downloaded. Only TNM stage I–III patients were enrolled. The datasets GSE42127,Citation21 GSE14814,Citation22 and GSE68465Citation23 were included (Tables S1 and S2).
Gene Set Enrichment Analysis
To investigate the potential molecular mechanisms during LARGE1 alteration in lung cancer, the Spearman correlations analysis results of more than 20,000 genes with LARGE1 in lung adenocarcinoma in the TCGA database were downloaded via cBioPortal (cbiopartal.org).Citation24,Citation25 The genes were ranked by the Spearman’s rank r and subjected to Gene Set Enrichment Analysis (GSEA) via software, JAVA GSEA 3.0 (http://software.broadinstitute.org/gsea/index.jsp).Citation26,Citation27 The biologically defined gene sets “c2.cp.kegg.v7.1.symbols.gmt,” “c7.all.v7.1.symbols.gmt,” and “h.all.v7.1.symbols.gmt” were used as the reference databases.
Association of LARGE mRNA Expression with Cell Cycle Pathway Score
Akbani et al have used reverse phase protein array (RPPA) data from TCGA to calculate the score for 10 cancer-related pathways in 32 cancer types including cell cycle pathway.Citation28 RPPA is a high-throughput antibody-based technique with procedures similar to that of Western blots. The pathway score is then the sum of the relative protein level of all positive regulatory components minus that of negative regulatory components in a particular pathway. For the cell cycle pathway, its score is the sum of the relative protein levels of CDK1, CYCLINB1, CYCLIND1, CYCLINDE1, CYCLINE2, P27PT157, P27PT198, and PCNA. Association of LARGE1 expression with cell cycle pathway score in lung adenocarcinoma was analyzed by Spearman correlation analysis.
Statistical Analysis
Kruskal–Wallis tests were used to compare LARGE1 expression in NSCLC tumor tissues, adjacent tissues, and normal lung tissues and in patients with different stages. Fisher’s exact test or Chi-square test were used to determine the associations of LARGE1 expression with clinical features of NSCLC patients. Kaplan-Meier curves and univariate Cox proportional hazards regression models as well as Kaplan-Meier curves adjusted by confounding factors and multivariate Cox proportional hazards regression models were used to clarify the effects of LARGE1 expression on OS in NSCLC patients. All statistical analyses were performed with R software (v 3.4.3). A two-sided P < 0.05 was considered statistically significant.
Results
LARGE1 Was Downregulated in NSCLC
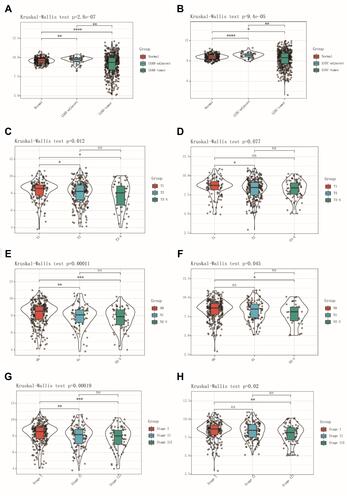
TCGA TARGET GTEx dataset was used to investigate LARGE1 mRNA expression in NSCLC tumor tissues, adjacent tissues, and normal lung tissues. LARGE1 was found to be downregulated in LUAD tumor tissues () and LUSC tumor tissues () compared with corresponding adjacent tissues and normal lung tissues. In addition, LARGE1 expression was determined in tumors with different stages in patients with TNM I–III NSCLC. The results suggested that LARGE1 was decreasingly expressed along with advancing tumor T stage ( and ), N stage ( and ), and TNM stage ( and ) in both LUAD (, , and ) and LUSC (, , and ), especially in LUAD.
Figure 1 LARGE1 mRNA expression in NSCLC. (A, B) LARGE1 expression in lung adenocarcinoma (LUAD) (A) and lung squamous cell carcinoma (LUSC) (B) compared to corresponding adjacent tissues and normal lung tissues. (C–H) LARGE1 expression in LUAD (C, E, and G) and LUSC (D, F, and H) with different T stages (C and D), N stages (E and F), and TNM stages (G and H).
Correlations of LARGE1 Expression with Clinicopathological Features in Stage I-III NSCLC Patients
A total of 981 NSCLC patients from the TCGA database with baseline clinical characteristics and OS outcome information were split into a high LARGE1 expression group and a low LARGE1 expression group by the median value of LARGE1 expression. As shown in , high LARGE1 was associated with advanced T stage, N stage, TNM stage, LUSC, and older age but not with gender and smoking history.
Table 1 Correlations of LARGE1 Expression with Clinicopathological Features in Stage I–III NSCLC Patients
Correlations of LARGE1 Expression Levels with Overall Survival in Stage I-III NSCLC Patients
Effects of LARGE1 expression levels on OS in NSCLC was first examined using Kaplan-Meier plotter (KM plotter) and high LARGE1 expression was found to be associated with favorable OS in 1,925 NSCLC patients (). Then the effect of LARGE1 expression on OS was investigated in the 981 NSCLC patients from TCGA. Increased LARGE1 expression was associated with favorable OS in NSCLC revealed by both Kaplan-Meier curve with Log rank test (optimal cutoff value, P=0.028; ) and univariate Cox proportional hazards regression model (HR=0.797; 95% CI=0.651–0.976; P=0.029). However, the association was not significant in the multivariate model (HR=0.806, 95% CI=0.633–1.027, P=0.081). Next, we used LARGE1 expression level as a continuous variable to explore its effects on OS in different subgroups stratified by age, gender, smoking history, histology, TNM stage, margin, adjuvant chemotherapy, and adjuvant radiotherapy. Association of increasing LARGE1 expression with OS was observed in the subgroup of patients receiving adjuvant chemotherapy but not in other subgroups (). After splitting LARGE1 expression into high and low groups, similar results were obtained by Kaplan-Meier curves and Cox regression analyses (; ). And ACT was found to provide more benefit in the high LARGE1 group compared to the low LARGE1 group ( and , and ). We also found a significant interaction between LARGE1 expression and adjuvant chemotherapy when evaluating their effects on OS (). Furthermore, we classified the patients by combining LARGE1 expression and adjuvant chemotherapy into three groups, ie, non-ACT, low LARGE1/ACT, and high LARGE1/ACT. Compared with patients with non-ACT, HR of low LARGE1/ACT was 0.592 (95% CI=0.432–0.813, P=0.0012), and HR of high LARGE1/ACT was 0.124 (95% CI=0.031–0.505, P=0.0036) ( and , and ).
Table 2 Correlations of LARGE1 Expression and Chemotherapy with Overall Survival in Different Subgroups of Stage I–III NSCLC Patients at Multivariate Analyses
Figure 2 Effects of LARGE1 on overall survival (OS) in NSCLC. (A) OS difference between NSCLC patients with high and low LARGE1 expression analyzed by the web-tool, Kaplan-Meier plotter, using auto select best cutoff value and the JetSet best probe set (215543_s_at). (B) OS difference between TNM I–III NSCLC patients with high and low LARGE1 expression from TCGA, analyzed by Kaplan-Meier curve with Log rank test. (C) Forest plot of the hazard ratios (HRs) for the associations of increasing LARGE1 expression (serves as a continuous variable) with overall survival (OS) of patients with TNM I–III NSCLC in subgroups stratified by age, gender, smoking history, histology, TNM stage, margin, adjuvant chemotherapy, and adjuvant radiotherapy.
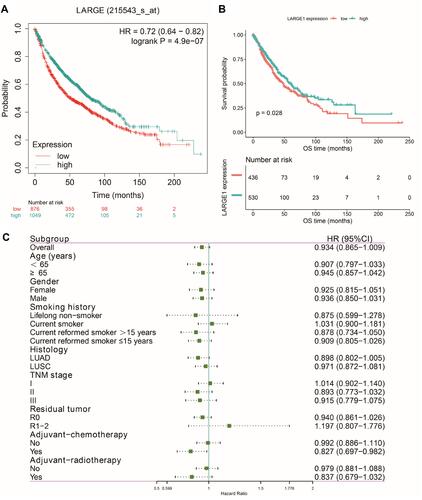
Figure 3 OS difference between TNM I–III NSCLC patients with high and low LARGE1 expression split by optimal cutoff value in subgroups stratified by adjuvant chemotherapy treatment (ACT) and between NSCLC patients with and without ACT in subgroups stratified by LARGE1 expression, demonstrated by Kaplan-Meier curves. (A) OS difference between NSCLC patients with high and low LARGE1 expression in the subgroup with ACT. (B) OS difference between NSCLC patients with high and low LARGE1 expression in subgroup of ACT adjusted by the confounding factors. (C) OS difference between NSCLC patients with high and low LARGE1 expression in subgroup of non-ACT adjusted by the confounding factors. (D) OS difference between NSCLC patients with and without ACT in the subgroup of low LARGE1 expression adjusted by the confounding factors. (E) OS difference between NSCLC patients with and without ACT in the subgroup of high LARGE1 expression adjusted by the confounding factors.
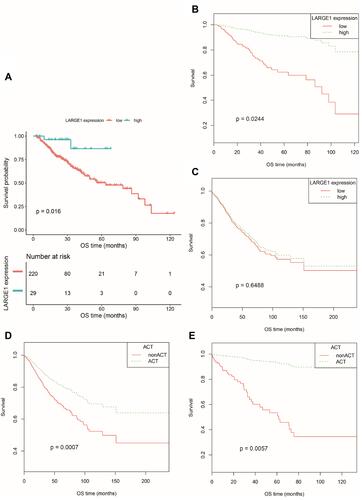
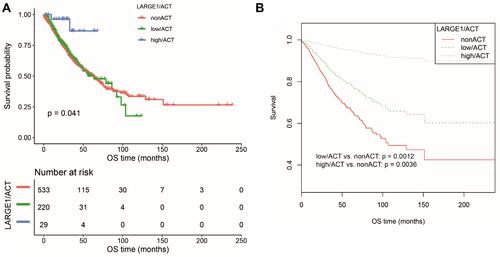
Figure 4 OS difference in TNM I–III NSCLC patients stratified by combining LARGE1 expression and adjuvant chemotherapy (ACT), ie, non-ACT, low LARGE1/ACT, and high LARGE1/ACT. (A and B) LARGE1 expression was split into high and low groups by optimal cutoff value; OS difference of patients with non-ACT, low LARGE1/ACT, and high LARGE1/ACT was analyzed by Kaplan-Meier curve (A), which was also analyzed by adjusting the confounding factors (B).
Validation
The above results suggested high LARGE1 expression might help to identify the patients who would get more benefit from adjuvant chemotherapy in stage I–III NSCLC, so next we validated the results in external cohorts from the GEO database. Datasets GSE42127, GSE14814, GSE29013, and GSE68465 contained resected NSCLC cases treated with ACT or not and provided OS information (Table S1). ACT was found to improve OS in datasets GSE42127 and GSE14814 while decrease OS in datasets GSE29013 and GSE68465, thus validation was only performed in datasets GSE42127 and GSE14814 (Table S2). Similar to the results from the TCGA cohort, in both GSE42127 and GSE14814 cohorts, LARGE1 expression was associated with OS in patients with ACT while not in patients without ACT, and ACT was associated with OS in patients with high LARGE1 expression while not in patients with low LARGE1 expression (Figures S1 and S2; ). The whole cohort of patients was also classified into three groups based on LARGE1 expression and ACT as well as TCGA cohort, compared to patients with non-ACT and those with LARGE1/ACT, patients with high LARGE1/ACT were found to be associated with significantly improved OS (Figures S3 and S4; ).
Potential Mechanisms During LARGE1 Alteration in Lung Adenocarcinoma
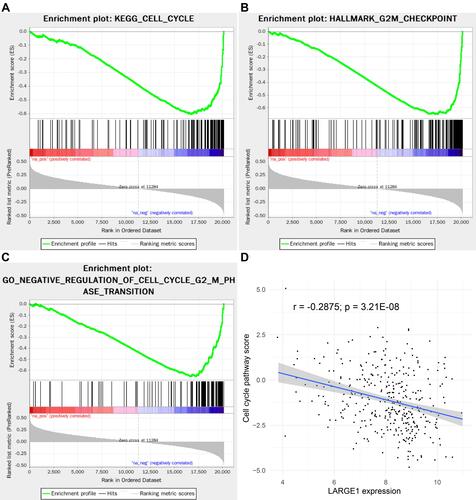
To elucidate the potential mechanisms during LARGE1 alteration in lung cancer, the Spearman correlations analysis results of more than 20,000 genes with LARGE1 in lung adenocarcinoma were download from TCGA. The genes were ranked by the Spearman’s rank r and subjected to Gene Set Enrichment Analysis (GSEA). KEGG pathways, GO biological processes, and hallmarks that were positively or negatively correlated with LARGE1 expression were explored. showed the top 10 negative- and top 10 positive-associated gene sets against KEGG (), HALLMARK (), and GO biological processes () databases, respectively. KEGG Cell cycle pathway (), HALLMARK G2/M checkpoint (), and GO cell cycle G2/M phase transition () were all negatively associated with LARGE1 expression, indicating LARGE1 might improve patients’ outcomes via partly downregulating cell cycle in lung adenocarcinoma. The result was further validated by examining the association of LARGE1 expression with cell cycle pathway score, which was calculated by summing cell cycle-related proteins expression ().
Figure 5 Gene Set Enrichment Analysis (GSEA) of LARGE1 co-expressed genes in lung adenocarcinoma (LUAD). Genes were ranked by Spearman’s rank rho association with LARGE1 expression were subjected to GSEA with KEGG pathways, HALLMARKS, and GO biological processes as reference databases. (A) The top 10 negative and top 10 positive KEGG pathways associated with LARGE1 expression. (B) The top 10 negative and top 10 positive HALLMARKS associated with LARGE1 expression. (C) The top 10 negative and top 10 positive GO biological processes associated with LARGE1 expression.
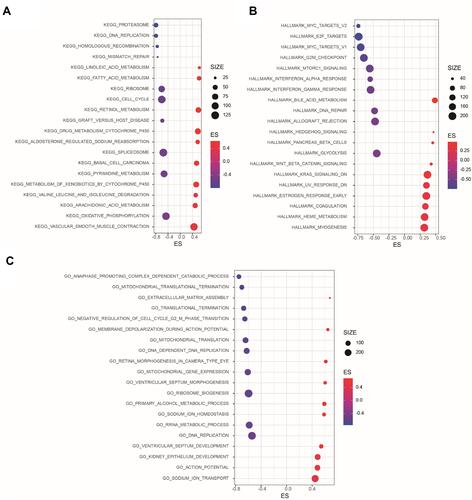
Figure 6 LARGE1 expression was negatively associated with cell cycle in lung adenocarcinoma (LUAD). (A–C) KEGG Cell cycle pathway (A), HALLMARK G2/M checkpoint (B), and GO cell cycle G2/M phase transition (C) were all negatively associated with LARGE1 expression. (D) The association of LARGE1 expression with cell cycle pathway score, which was calculated by summing cell cycle-related proteins expression.
Discussion
α-Dystroglycan functions as an extracellular matrix (ECM) receptorCitation29 for several LG domain-containing basement membrane proteins via a unique heteropolysaccharide [-GlcA-beta1,3-Xyl-alpha1,3-]n called matriglycan, including laminin,Citation30–Citation32 perlecan,Citation33 agrin,Citation34 and Slit.Citation35 α-DG has been implicated in several cell functions (ie, growth control, differentiation, shape change and movement) which are all relevant in the process of tumor development and metastasis.Citation36,Citation37 Expression or glycosylation of α-DG has been identified to be lost or reduced in a variety of human cancer cell linesCitation17,Citation38 and different cancers including rhabdomyosarcoma,Citation18 gliomas,Citation39 breast,Citation40,Citation41 prostate,Citation42 colon,Citation41,Citation43 cervical,Citation44 vulvar,Citation44 gastric,Citation45 tongue,Citation46 and pancreatic cancers,Citation47 oral squamous cell carcinoma,Citation48 and renal cell carcinoma.Citation49 Such reduction or loss of α-DG seems to correlate with the biologically invasive character of primary tumorsCitation41,Citation44,Citation46,Citation47,Citation49–Citation51 and associated with poor outcomes.Citation41,Citation43,Citation45,Citation47,Citation49,Citation51 LARGE1 is important for α-DG glycosylation by adding the laminin-binding repeating units of 3GlcA-1-3Xyl-1 onto α-DGCitation16 and significant association of LARGE1 downregulation with abnormal α-DG glycosylation in cancer cells has been observed.Citation17,Citation18 In the present study, we, for the first time, have reported that LARGE1 mRNA expression was downregulated in lung cancer tissues compared with adjacent tissues and normal lung tissues and in patients with advanced stages compared with early stages via analyzing the large dataset from the TCGA database. And there was a trend of significant association between LARGE1 downregulation with poor overall survival in stage I–III patients in overall and LARGE1 downregulation was significantly associated with poor overall survival in stage I–III patients receiving ACT.
ACT is the standard care for patients with NSCLC after complete resection, unfortunately, only a few of the patients get to benefit from ACT due to primary and acquired drug resistance.Citation14,Citation15 Therefore, it is critical to develop predictive biomarkers to identify the patients who will benefit from ACT and who will not benefit from ACT to avoid unnecessary toxicity and the delay of further treatment. Previously, low expression of ERCC1Citation52 and high expression of TUBB3Citation53 were shown to be predictive of benefit from ACT in NSCLC patients from a large cohort or clinical trial. However, large cross-validation studies suggested that attempts to validate ERCC1 or TUBB3 as predictive biomarkers fell short.Citation54,Citation55 Similar results were derived for validating other single biomarkers including p53, RRM1, and p27.Citation56–Citation58 Here, we found high LARGE1 mRNA expression can be used to identify the patients who would benefit from ACT, which was also validated in external cohorts, although this should be validated in more and larger multi-center studies.
LARGE1 is essential for α-DG glycosylation, which mediates the interaction between cells and the ECM and may play critical roles in the development and progression of many cancers by modulating cell growth, differentiation, shape change, and movement.Citation36,Citation37,Citation59 In our study, we performed GSEA and found cell cycle regulation during LARGE1 alteration might be involved in carcinogenesis of lung adenocarcinoma, which might also be associated with ACT sensitivity. The potential mechanisms of LARGE1 downregulation in lung cancer were also unknown. Zhang et al reported that reduced LARGE1 expression was associated with its promoter hypermethylation in tongue cancer.Citation46 In lung cancer, we also identified the negative and significant association between LARGE1 mRNA expression and the methylation of the CpG islands at its promoter (data not shown). More in vitro and animal experiments should be performed to validate our results.
Conclusions
LARGE1 mRNA expression was aberrantly downregulated in NSCLC and especially in cancers with advanced stages. High LARGE1 mRNA expression was associated with favorable OS in NSLCLC patients undergoing adjuvant chemotherapy rather than in patients without ACT and improved OS after ACT was also found in patients with high LARGE1 than those with low LARGE1. External cohorts confirmed these results, indicating that high LARGE expression could be used to identify the NSCLC patients who benefited from adjuvant chemotherapy.
Acknowledgment
This study was supported by the subjects from the Natural Science Foundation of China (NSFC81601849), Zhejiang Provincial Medicine and Health Technology Project (2019RC217), Wenzhou Science and Technology Bureau (Y20180109, Y20180254).
Disclosure
The authors report no conflicts of interest in this work.
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):425.
- Travis WD. Pathology of lung cancer. Clin Chest Med. 2011;32(4):669–692. doi:10.1016/j.ccm.2011.08.005
- Torre LA, Siegel RL, Jemal A. Lung cancer statistics. Adv Exp Med Biol. 2016;893:893. doi:10.1007/978-3-319-24223-1_1
- Xue C, Hu Z, Jiang W, et al. National survey of the medical treatment status for non-small cell lung cancer (NSCLC) in China. Lung Cancer. 2012;77(2):371–375. doi:10.1016/j.lungcan.2012.04.014
- Hung -J-J, Jeng W-J, Hsu W-H, Huang B-S, Wu Y-C. Time trends of overall survival and survival after recurrence in completely resected stage I non-small cell lung cancer. J Thorac Oncol. 2012;7(2):397–405. doi:10.1097/JTO.0b013e31823b564a
- Su S, Scott WJ, Allen MS, et al. Patterns of survival and recurrence after surgical treatment of early stage non–small cell lung carcinoma in the ACOSOG Z0030 (ALLIANCE) trial. J Thorac Cardiovasc Surg. 2014;147(2):2. doi:10.1016/j.jtcvs.2013.10.001
- Chen W, Zheng R, Zhang S, Zhao P, Zeng H, Zou X. Report of cancer incidence and mortality in China, 2010. Ann Transl Med. 2014;2(7):61. doi:10.3978/j.issn.2305-5839.2014.04.05
- Galluzzi L, Buqué A, Kepp O, Zitvogel L, Kroemer G. Immunological effects of conventional chemotherapy and targeted anticancer agents. Cancer Cell. 2015;28(6):690–714. doi:10.1016/j.ccell.2015.10.012
- Arriagada R, Bergman B, Dunant A, Le Chevalier T. Cisplatin-based adjuvant chemotherapy in patients with completely resected non-small-cell lung cancer. N Engl J Med. 2004;350(4):351–360.
- Waller D, Peake MD, Stephens RJ, et al. Chemotherapy for patients with non-small cell lung cancer: the surgical setting of the Big Lung Trial. Eur J Cardiothorac Surg. 2004;26(1):173–182. doi:10.1016/j.ejcts.2004.03.041
- Winton T, Livingston R, Johnson D, et al. Vinorelbine plus cisplatin vs. observation in resected non–small-cell lung cancer. N Engl J Med. 2005;352(25):2589–2597. doi:10.1056/NEJMoa043623
- Douillard J-Y, Rosell R, De Lena M, et al. Adjuvant vinorelbine plus cisplatin versus observation in patients with completely resected stage IB–IIIA non-small-cell lung cancer (Adjuvant Navelbine International Trialist Association [ANITA]): a randomised controlled trial. Lancet Oncol. 2006;7(9):719–727. doi:10.1016/S1470-2045(06)70804-X
- Strauss GM, Herndon JE, Maddaus MA, et al. Adjuvant paclitaxel plus carboplatin compared with observation in stage IB non–small-cell lung cancer: CALGB 9633 with the Cancer and Leukemia Group B, Radiation Therapy Oncology Group, and North Central Cancer Treatment Group Study Groups. J Clin Oncol. 2008;26(31):5043–5051. doi:10.1200/JCO.2008.16.4855
- Pignon J-P, Tribodet H, Scagliotti GV, et al. Lung adjuvant cisplatin evaluation: a pooled analysis by the LACE Collaborative Group. J Clin Oncol. 2008;26(21):3552–3559. doi:10.1200/JCO.2007.13.9030
- Kelland L. The resurgence of platinum-based cancer chemotherapy. Nat Rev Cancer. 2007;7(8):573–584. doi:10.1038/nrc2167
- Inamori K, Hara Y, Willer T, et al. Xylosyl- and glucuronyltransferase functions of LARGE in α-dystroglycan modification are conserved in LARGE2. Glycobiology. 2013;23(3):295–302. doi:10.1093/glycob/cws152
- de Bernabé DB-V, Inamori K-I, Yoshida-Moriguchi T, et al. Loss of α-dystroglycan laminin binding in epithelium-derived cancers is caused by silencing of LARGE. J Biol Chem. 2009;284(17):11279–11284. doi:10.1074/jbc.C900007200
- Beltrán D, Anderson ME, Bharathy N, et al. Exogenous expression of the glycosyltransferase LARGE1 restores α-dystroglycan matriglycan and laminin binding in rhabdomyosarcoma. Skeletal Muscle. 2019;9(1):11. doi:10.1186/s13395-019-0195-0
- Goldman MJ, Craft B, Hastie M, et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat Biotechnol. 2020;38(6):675–678. doi:10.1038/s41587-020-0546-8
- Győrffy B, Surowiak P, Budczies J, Lánczky A, Chellappan SP. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS One. 2013;8(12):e82241. doi:10.1371/journal.pone.0082241
- Tang H, Xiao G, Behrens C, et al. A 12-Gene Set Predicts Survival Benefits from Adjuvant Chemotherapy in Non–Small Cell Lung Cancer Patients. Clin Cancer Res. 2013;19(6):1577–1586. doi:10.1158/1078-0432.CCR-12-2321
- Zhu C-Q, Ding K, Strumpf D, Zhu CQ, Ding K, Strumpf D, et al. Prognostic and Predictive Gene Signature for Adjuvant Chemotherapy in Resected Non–Small-Cell Lung Cancer. J Clin Oncol. 2010;28(29):4417–4424. doi:10.1200/JCO.2009.26.4325
- Lung A, Shedden K, Taylor JM, et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med. 2008;14(8):822–827.
- Cerami E, Gao J, Dogrusoz U, et al. The cBio Cancer Genomics Portal: an open platform for exploring multidimensional cancer genomics data: figure 1. Cancer Discov. 2012;2(5):401. doi:10.1158/2159-8290.CD-12-0095
- Gao J, Aksoy BA, Dogrusoz U, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):pl1. doi:10.1126/scisignal.2004088
- Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545. doi:10.1073/pnas.0506580102
- Mootha VK, Lindgren CM, Eriksson K-F, et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34(3):267–273. doi:10.1038/ng1180
- Akbani R, Ng PKS, Werner HMJ, et al. A pan-cancer proteomic perspective on The Cancer Genome Atlas. Nat Commun. 2014;5(1):3887. doi:10.1038/ncomms4887
- Barresi R. Dystroglycan: from biosynthesis to pathogenesis of human disease. J Cell Sci. 2006;119(2):199–207. doi:10.1242/jcs.02814
- Yamada H, Denzer AJ, Hori H, et al. Dystroglycan is a dual receptor for agrin and laminin-2 in Schwann cell membrane. J Biol Chem. 1996;271(38):23418–23423. doi:10.1074/jbc.271.38.23418
- Kanagawa M, Saito F, Kunz S, et al. Molecular recognition by LARGE is essential for expression of functional dystroglycan. Cell. 2004;117(7):953–964. doi:10.1016/j.cell.2004.06.003
- Kunz S, Sevilla N, McGavern DB, Campbell KP, Oldstone MBA. Molecular analysis of the interaction of LCMV with its cellular receptor α-dystroglycan. J Cell Biol. 2001;155(2):301–310. doi:10.1083/jcb.200104103
- Gee SH, Montanaro F, Lindenbaum MH, Carbonetto S. Dystroglycan-α, a dystrophin-associated glycoprotein, is a functional agrin receptor. Cell. 1994;77(5):675–686. doi:10.1016/0092-8674(94)90052-3
- Talts JF, Andac Z, Göhring W, Brancaccio A, Timpl R. Binding of the G domains of laminin α1 and α2 chains and perlecan to heparin, sulfatides, α-dystroglycan and several extracellular matrix proteins. EMBO J. 1999;18(4):863–870. doi:10.1093/emboj/18.4.863
- Wright KM, Lyon KA, Leung H, Leahy DJ, Ma L, Ginty DD. Dystroglycan organizes axon guidance cue localization and axonal pathfinding. Neuron. 2012;76(5):931–944. doi:10.1016/j.neuron.2012.10.009
- Sgambato A, Brancaccio A. The dystroglycan complex: from biology to cancer. J Cell Physiol. 2005;205(2):163–169. doi:10.1002/jcp.20411
- Sgambato A, Di Salvatore MA, De Paola B, et al. Analysis of dystroglycan regulation and functions in mouse mammary epithelial cells and implications for mammary tumorigenesis. J Cell Physiol. 2006;207(2):520–529. doi:10.1002/jcp.20600
- Losasso C, Di Tommaso F, Sgambato A, et al. Anomalous dystroglycan in carcinoma cell lines. FEBS Lett. 2000;484(3):194–198. doi:10.1016/S0014-5793(00)02157-8
- Calogero A, Pavoni E, Gramaglia T, et al. Altered expression of alpha-dystroglycan subunit in human gliomas. Cancer Biol Ther. 2006;5(4):441–448. doi:10.4161/cbt.5.4.2546
- Sgambato A, Camerini A, Faraglia B, et al. Increased expression of dystroglycan inhibits the growth and tumorigenicity of human mammary epithelial cells. Cancer Biol Ther. 2004;3(10):967–975. doi:10.4161/cbt.3.10.1132
- Sgambato A, Migaldi M, Montanari M, et al. Dystroglycan expression is frequently reduced in human breast and colon cancers and is associated with tumor progression. Am J Pathol. 2003;162(3):849–860. doi:10.1016/S0002-9440(10)63881-3
- Sgambato A, De Paola B, Migaldi M, et al. Dystroglycan expression is reduced during prostate tumorigenesis and is regulated by androgens in prostate cancer cells. J Cell Physiol. 2007;213(2):528–539. doi:10.1002/jcp.21130
- Coco C, Zannoni GF, Caredda E, et al. Increased expression of CD133 and reduced dystroglycan expression are strong predictors of poor outcome in colon cancer patients. J Exp Clin Cancer Res. 2012;31(1):71. doi:10.1186/1756-9966-31-71
- Sgambato A, Tarquini E, Resci F, et al. Aberrant expression of α-dystroglycan in cervical and vulvar cancer. Gynecol Oncol. 2006;103(2):397–404. doi:10.1016/j.ygyno.2006.03.059
- Moon YW, Rha SY, Zhang X, et al. Increments of α-dystroglycan expression in liver metastasis correlate with poor survival in gastric cancer. J Surg Oncol. 2009;100(6):459–465. doi:10.1002/jso.21347
- Zhang H-Z, Xia X-Y, Zhu F, Shen H, Song K, Shang Z-J. Correlation of deregulated like-acetylglucosaminyl transferase and aberrant α-dystroglycan expression with human tongue cancer metastasis. J Oral Maxillofac Surg. 2014;72(6):1106–1118. doi:10.1016/j.joms.2013.12.031
- Jiang X, Rieder S, Giese NA, Friess H, Michalski CW, Kleeff KJ. Reduced α-dystroglycan expression correlates with shortened patient survival in pancreatic cancer. J Surg Res. 2011;171(1):120–126. doi:10.1016/j.jss.2009.11.730
- Jing J, Lien CF, Sharma S, Rice J, Brennan PA, Górecki DC. Aberrant expression, processing and degradation of dystroglycan in squamous cell carcinomas. Eur J Cancer. 2004;40(14):2143–2151. doi:10.1016/j.ejca.2004.05.018
- Sgambato A, Camerini A, Amoroso D, et al. Expression of dystroglycan correlates with tumor grade and predicts survival in renal cell carcinoma. Cancer Biol Ther. 2007;6(12):1840–1846. doi:10.4161/cbt.6.12.4983
- Henry MD, Cohen MB, Campbell KP. Reduced expression of dystroglycan in breast and prostate cancer. Hum Pathol. 2001;32(8):791–795. doi:10.1053/hupa.2001.26468
- Shen JG, Xu CY, Li X, et al. Dystroglycan is associated with tumor progression and patient survival in gastric cancer. Pathol Oncol Res. 2012;18(1):79–84. doi:10.1007/s12253-011-9419-2
- Olaussen KA, Dunant A, Fouret P, et al. DNA repair by ERCC1 in non–small-cell lung cancer and cisplatin-based adjuvant chemotherapy. N Engl J Med. 2006;355(10):983–991. doi:10.1056/NEJMoa060570
- Sève P, Lai R, Ding K, Sève P, Lai R, Ding K, et al. Class III β-tubulin expression and benefit from adjuvant cisplatin/vinorelbine chemotherapy in operable non–small cell lung cancer: analysis of NCIC JBR.10. Clin Cancer Res. 2007;13(3):994–999. doi:10.1158/1078-0432.CCR-06-1503
- Reiman T, Lai R, Veillard AS, et al. Cross-validation study of class III beta-tubulin as a predictive marker for benefit from adjuvant chemotherapy in resected non-small-cell lung cancer: analysis of four randomized trials. Ann Oncol. 2012;23(1):86–93. doi:10.1093/annonc/mdr033
- Friboulet L, Olaussen KA, Pignon J-P, et al. ERCC1 isoform expression and DNA repair in non–small-cell lung cancer. N Engl J Med. 2013;368(12):1101–1110. doi:10.1056/NEJMoa1214271
- Olaussen KA, Postel-Vinay S. Predictors of chemotherapy efficacy in non-small-cell lung cancer: a challenging landscape. Ann Oncol. 2016;27(11):2004–2016. doi:10.1093/annonc/mdw321
- Wallerek S, Sørensen JB. Biomarkers for efficacy of adjuvant chemotherapy following complete resection in NSCLC stages I–IIIA. Eur Respir Rev. 2015;24(136):340–355. doi:10.1183/16000617.00005814
- Seymour L, Le Teuff G, Brambilla E, et al. LACE-Bio: validation of predictive and/or prognostic immunohistochemistry/histochemistry-based biomarkers in resected non–small-cell lung cancer. Clin Lung Cancer. 2019;20(2):2. doi:10.1016/j.cllc.2018.10.001
- Petersen OW, Rønnov-Jessen L, Howlett AR, Bissell MJ. Interaction with basement membrane serves to rapidly distinguish growth and differentiation pattern of normal and malignant human breast epithelial cells. Proc Natl Acad Sci U S A. 1992;89(19):9064–9068. doi:10.1073/pnas.89.19.9064
