Abstract
Pyrococcales are members of the order Thermococcales, a group of hyperthermophilic euryarchaea that are frequently found in deep sea hydrothermal vents. Infectious genetic elements, such as plasmids and viruses, remain a threat even in this remote environment and these microorganisms have developed several ways to fight their genetic invaders. Among these are the recently discovered CRISPR systems. In this review, we have combined and condensed available information on genetic elements infecting the Thermococcales and on the multiple CRISPR systems found in the Pyrococcales to fight them. Their organization and mode of action will be presented with emphasis on the Type III-B system that is the only CRISPR system known to target RNA molecules in a process reminiscent of RNA interference. The intriguing case of Pyrococcus abyssi, which is among the rare strains to present a CRISPR system devoid of the universal cas1 and cas2 genes, is also discussed.
Introduction
Pyrococcus species belong to the Thermococcales, which are one of the predominant groups of microorganisms isolated from the deep sea hydrothermal vents. Similar to organisms in any other ecological habitats, these microorganisms face constant attacks from invading genetic elements such as phages and plasmids. Among the multiple defense systems that have been developed to counteract infectious genetic elements in bacteria and archaea, one is of particular interest: the CRISPR (clustered regularly interspaced short palindromic repeats) systems, which are found in 90% of known archaeal genomes and in about half of the known bacterial genomes (CRISPRdb databaseCitation1). It was recently discovered that CRISPR loci function as adaptive and inheritable immunity systems, which are responsible for defending the cell against invading viruses and plasmids (for recent reviews see refs.Citation2–Citation5). CRISPR loci are composed of arrays of direct repeats separated by variable sequences called spacers or guides that are generally derived from invader genetic elements. A match between a CRISPR guide and an invading nucleic acid provides immunity to infection. Within a locus, repeat and guide sequences have conserved sizes, which can range between 23–47 base pairs depending on the system.
A dozen major groups of cas (CRISPR-associated) gene families,Citation6 located near CRISPR loci in various organisms, are proposed to be involved in the three major phases of the CRISPR systems, which are the adaptation, expression and interference phases. During adaptation, protospacer sequences (foreign DNA sequences acquired into the host genome) are selected among invader DNA and inserted at the leader (region upstream of a CRISPR array that contains a promoter) proximal end of CRISPR arrays to generate additional guide sequences. The expression phase consists of the transcription of the CRISPR arrays to produce long pre-crRNA (pre-CRISPR RNA), which are further cleaved at each repeats into individual crRNA. Finally, during the interference phase, invading nucleic acids that match a crRNA are restricted by effector ribonucleoprotein (RNP) complexes consisting of dedicated Cas proteins associated to individual crRNAs. The proteins encoded by the cas genes include predicted RNA-binding proteins, endo- and exo-nucleases, helicases and polymerases (for a review see ref. Citation6). Most of these cas genes can be clustered into three major CRISPR/cas system types (Type I, II and III), which are further divided into 10 subtypes (subtypes I-A to -F, subtypes II-A and -B and subtypes III-A and -B).Citation7 The conserved “core” of CRISPR systems is composed of the six cas1 to cas6 genes but the only universal CRISPR genes seem to be cas1 and cas2, which are found in almost every organism bearing a CRISPR system (e.g., Pyrococcus abyssi lacks cas1 and cas2). Multiple CRISPR/cas system types can coexist in a single genome and a set of cas genes can be functionally linked to several CRISPR loci.Citation6
Only few infectious genetic elements have been identified so far in Thermococcales, whereas they have all been found to contain elements of CRISPR/cas systems in their genomes. CRISPR/cas systems of Thermococcus species are still partially annotated, particularly the cas genes, due to the absence of available experimental data. Conversely, pyrococcal CRISPR/cas systems have been more accurately annotated based on the significant biochemical information on crRNA processing and Cas RNP complexes. Thus, this review will mention the infectious genetic elements found within the Thermococcales and will emphasize the diversity of CRISPR/cas systems only in Pyrococcales since information in Thermococcus species are still sparse. Information on the pyrococcal CRISPR/cas systems was combined and summarized, using the six available pyrococcal genomes to illustrate the complexity and dynamics of CRISPR systems. The CRISPR/cas nomenclature and terminology proposed by Makarova and coworkers,Citation7 which associates information from phylogenetic and comparative analyses, is used throughout this review.
The Thermococcales and Their Infectious Genetic Elements
While archaea can be found in virtually every ecosystem, they tend to dominate in extreme environments presenting high salt concentrations, low pH or high temperatures. Pyrococcales are members of the order Thermococcales, which belong to the domain Euryarchaea and include also the genera Palaeococcus and Thermococcus. They are anaerobic chemoorganotrophs (heterotrophs) and necessitate rich media containing peptides and organic compounds for optimal growth. These organic compounds are oxidized to provide electrons that are ultimately transferred to sulfur to form H2S. Typically, Thermococcales are small cocci with polar flagella that provide good motility. They have been identified in diverse hot aquatic ecological habitats, including oceanic and fresh waterCitation8 hydrothermal vents, oil reservoirsCitation9,Citation10 and even subsurface environments.Citation11,Citation12 Their optimum growing temperatures range from 70–106°C. Due to their high diversity (more than 40 reported species) and abundance in hot aquatic ecosystems, Thermococcales are studied in many laboratories as model organisms for hyperthermophiles.Citation13 Thermococcales and other hyperthermophiles are industrially important because they are source of thermostable biocatalysts permitting the development of high temperature biotechnological applications.
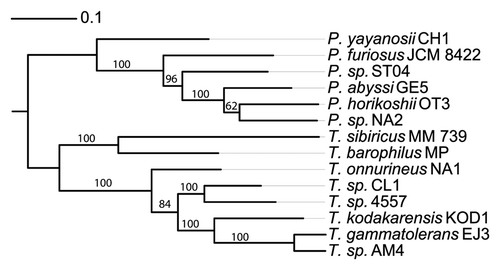
Genome sequences from 14 Thermococcales are now available, including six genomes of Pyrococcus (P. abyssi GE5;Citation14 P. horikoshii;Citation15 P. furious;Citation16; P. yayanosii;Citation17; Pyrococcus sp NA2;Citation18 Pyrococcus sp ST04Citation19) and eight genomes of Thermococcus (T. kodakarensis KOD1;Citation20 T. onnurineus NA1;Citation20 T. barophilus MP;Citation20 T. sibiricus MM739;Citation9 T. gammatolerans EJ3;Citation21 Thermococcus sp AM4;Citation22 Thermococcus sp 4557;Citation23 Thermococcus CL1Citation24). A phylogenetic tree of these Thermococcales is shown in . The availability of these sequences makes Thermococcales a good model to explore the hyperthermophilic euryarchaeal virome, which remains poorly characterized. Comparative genomics among Thermococcales revealed important chromosome shuffling mediated by mobile genetics elements.Citation25 These genomes are highly divergent and present little synteny, which is restricted to few operons.
Figure 1. Thermococcal phylogenetic tree. This tree is based on a set of concatenated sequence alignment of 252 groups of orthologous proteins conserved in the 14 genomes (Y. Quentin, unpublished data). According to a previous analysis,Citation76 the tree was rooted between Pyrococcus and Thermococcus clades.
Archaeal viruses present an astounding morphological diversity compared with known viruses from bacteria and eukaryotes.Citation26,Citation27 All but two archaeal viruses described thus far contain dsDNA genomes. The haloarchaeal virus HRPV1 and the Aeropyrum coil-shaped virus (ACV) being two exceptions with an ssDNA genome.Citation28,Citation29 Most of our knowledge concerning these infectious genetic elements in hyperthermophiles comes from studies conducted on thermoacidophilic crenarchaea of the order Sulfolobales.Citation26 A large spectrum of genetic elements, including cryptic, conjugative or integrative plasmids and lysogenic or lytic phages representing at least seven novel families of virus, has been described to infect Sulfolobales.Citation26 This contrasts with the paucity of information available concerning infectious genetic elements from hyperthermophilic euryarchaea. Despite a large number of described strains, very few plasmids and viruses are described in Thermococcales. This is surprising as the prevalence of such elements is reported to be high among the Thermococcales; 20% to half of them carry at least one extrachromosomal element ranging in size from 3–35 kb.Citation30,Citation31 Known plasmids from Thermococcales include pGT5Citation32 (3.4 kb) from Pyrococcus abyssi GE5, pAMT11Citation33 (20.5 kb) from Thermococcus species, pRT1Citation34 (3.4 kb) from Pyrococcus sp JT1, pTN1Citation35 (3.6 kb) and pTN2Citation36 (13 kb) from Thermococcus nautilus, pP12-1Citation36 (12 kb) from Pyrococcus sp 12-1 and pT26-2Citation36 (21.5 kb) from Thermococcus sp 26-2. Recently, five additional plasmids (pCIR10, pIRI48, pAMT7, pEXT9a and pIRI33) have been described from various Thermococcus species.Citation37
Only two viruses from Thermococcales have been identified so far. The first one, PAV1Citation38,Citation39 (18 kb), is found in Pyrococcus abyssi GE23. The second virus, TPV1Citation40 (21.5 kb) has been isolated from Thermococcus prieurii. Similar to the majority of known archeoviruses, PAV1 and TPV1 establish chronic infections where virions are released continuously in the extracellular medium, and the host remains alive (no cell lysis upon egress). Thus, PAV1 and TPV1 persist in their host strains in a stable carrier state. No insertion in the Pyrococcus abyssi GE23 genome could be detected for PAV1, which maintains itself as an episome with about 60 copies per cell.Citation39 In addition to being maintained as an episome (20 copies per cell), TPV1 might be able to integrate to T. prieurii genome.Citation40 Despite numerous attempts, no host other than P. abyssi GE23 could be infected by PAV1 particles. On the other end, TPV1 particles could infect numerous related Thermococcus strains, which makes this virus a promising tool for Thermococcales molecular genetics studies.Citation40 In addition, a number of putative integrated viruses have been identified in Thermococcales genomes: TKV1 to TKV4 in T. kodakarensis,Citation41 TGV1 and TGV2 in T. gammatoleransCitation21 and PHV1 in P. horikoshii.Citation36 The integration pattern of these elements is characterized by flanking regions constituted by the partitioned N-terminal and C-terminal fragments of an integrase gene related to SSV1.Citation42 Regions coding the N-terminal fragment of the integrase overlap with the 5′ region of genes encoding a tRNA-Val, tRNA-Glu or tRNA-Arg. These tRNA genes are predicted to form attachment sites (att) for integration of transposable elements.
Despite their extreme way of life, Thermococcales still have to face, like any other cell, numerous threats from infectious genetic elements. The CRISPR systems are proposed to be among the defense systems against genetic invaders. The rest of this review will present the organizations and modes of action of CRISPR systems from the Pyrococcales.
Organization of CRISPR/cas Loci in the Pyrococcales
Available information concerning pyrococcal CRISPR/cas systems was extracted from public databases such as GenBank,Citation43 UCSC archaeal genome browser,Citation44 TIGRFAMCitation45 and CRISPRdb,Citation1 and condensed to generate with the use of Makarova nomenclature.Citation7 Several CRISPR/cas system types appear in Pyrococcal genomes as already observed in other archaeal genomes.Citation46-Citation48 This illustrates once again the diversity, complexity and dynamism of CRISPR/cas systems. Remarkably, P. abyssi and P. species NA2 genomes encompass two times less CRISPR loci and cas-related genes than the other four Pyrococcales in which seven to eight CRISPR arrays and two to three cas gene clusters are annotated (). Intriguingly, P. species NA2 genome possesses a high number of repeats within its CRISPR arrays (up to 79 and 85 repeats in CRISPR 2 and CRISPR 3, respectively). Several CRISPR remnants with highly degenerated direct repeat sequences and a low number of guides are sporadically found in pyrococcal genomes (unpublished data). They will not be discussed here and are not mentioned in .
Figure 2. Genomic arrangements of CRISPR arrays and cas modules within pyrococcal genomes. The nomenclature and classification is based on the polythetic classification proposed by Makarova and collaborators.Citation7 The pyrococcal subtype IA system is organized in two distinct effector modules denoted I-A1 and I-A2 modules. Note that the subtype I-A system is defined upon the presence of the cas8a1/cst1 and cas8a2/csx9/csa4 signature genes as referred in Makarova et al. and in Haft et al.Citation7,Citation49 The CRISPR arrays extracted from CRISPRdbCitation1 in each genome are denoted by long arrows with guide sequences in green. Note that some of the CRISPR arrays are reoriented to preserve the most conserved direct repeat proximal to the leader sequence.Citation53 Group 1 direct repeats (DRs) associated to the subtype I-A system are represented in orange, whereas the Group 2 DRs connected to the subtype III-A system are in pink (see ). The number of their repeats is indicated in parenthesis. Effector cas genes are represented in yellow for subtype I-A, red for subtype III-B and purple for subtype III-A. The csx1 genes that encode putative transcriptional regulators are indicated in green. The cas1, cas2 and cas4 genes forming the informational module involved in the CRISPR adaptation phase are in blue. The cas6 genes, encoding a processing factor of pre-crRNA are in cyan. Additional uncharacterized open reading frames embedded in cas loci are indicated by white triangles (ND, not determined). Note that this schematic representation is not to scale.

Three different effector cas types, defined by the presence of specific signature genes,Citation7 were identified within the six pyrococcal genomes: subtype I-A, subtype III-A and subtype III-B systems, which code for the respective effector RNP complexes that are implicated in the interference phase (). Notably, the CRISPR/cas subtype I-A shows an organization in two modules, which are here delineated as modules I-A1 and I-A2, to highlight its versatility. The I-A1 module (cas6, cas8a1, cas7, cas5, cas3) and I-A2 module (csa5, cas7, cas5, cas8a2, cas3′, cas3”) are characterized by the presence of cas8a1/cst1 or cas8a2/csx9,csa4 signature genes,Citation7,Citation49 respectively. These two modules are co-oriented and not always clustered together within the same locus. The I-A2 modules are found in all pyrococcal genomes but rarely in direct proximity of a CRISPR array. Remarkably, this I-A2 module is the sole CRISPR/cas signature in P. abyssi, which could suggest that it constitutes a minimal core of subtype I-A system, or that P. abyssi, which also lacks cas1 and cas2, may have lost a subset of its cas genes. Interestingly, the I-A1 modules are absent in the P. species ST04 and P. abyssi genomes and are found relocated together with genes belonging to the subtype III-B system in P. furiosus genome. Thus, the I-A1 modules appear to be adaptable within pyrococcal genomes. On the other hand, subtype III-A and III-B systems are less common in pyrococcal genomes. P. yayanosii, P. horikoshii and P. species ST04 all carry a subtype III-A system but subtype III-B systems are only present in P. yayanosii and P. furiosus genomes. Interestingly, P. yayanosii genome is the sole genome to contain all three cas gene subtypes (I-A, III-A and III-B) that are delineated in three distinct clusters. Finally, the P. furiosus I-A1 module is found adjacent and co-oriented with a subtype III-B gene locus. This generates a hybrid I-A/III-B system, which is not unique to P. furiosus as hybrid systems are found in several other archaeal and bacterial genomes.Citation6,Citation7 This hybrid system suggests a cross-talk between the subtype I-A system that targets DNA and the III-B system that restricts RNA (as presented below).
The highly conserved cas1, cas2 and cas4 genes form the “informational” module involved in the adaptation phase,Citation6 known to be implicated in the acquisition of new spacers. These genes are located adjacent to a CRISPR array and generally clustered together in all pyrococcal genomes except for P. abyssi that appears to lack cas1 and cas2. The cas1, cas2 and cas4 genes belong to the set of CRISPR “core” genes and are predicted to encode, respectively, a metal-dependent nuclease, an endoribonuclease and a RecB-like nuclease.Citation6 The pyrococcal adaptation subsystems show two alternative organizations depending on the gene locus subtype with which they are associated (). Those adjacent to I-A1 module cas genes are characterized by an inverted cas2 gene within the cas2-cas1-cas4 cluster, and is located just upstream of a CRISPR array in the opposite direction, except in the P. yayanosii subtype I-A system where cas1 and cas4 are situated further away, between the I-A1 and I-A2 modules. Conversely the adaptation modules associated with subtype III-A cas genes are composed of the three successive and co-oriented cas1, cas2, and cas4 genes. In this configuration, the cas1 gene is adjacent to the CRISPR array. These two separate organizations may reflect a distinct function or co-regulated expression of these three core genes. Furthermore, subtype III-B system is not directly associated to any adaptation module in pyrococcal genomes. In addition to the adaptation modules, and reminiscent of the cas6 genes organization (see below), all genomes carry an individual cas4 gene, which is isolated from any cas gene cluster or CRISPR array ().
Two to three cas6 genes can be found in each pyrococcal genome. The P. furiosus PF1131 cas6 gene located at the beginning of the hybrid subtype I-A/III-B cluster encodes the endoribonuclease in charge of crRNA processingCitation50 (see below). The function of the second P. furiosus cas6 gene (cas6-2 in ref. Citation51), which is not associated with any other cas gene cluster, is to our knowledge uncharacterized. In addition, three groups of pyrococcal cas6 genes have been delineated as a result of sequence comparisons (Moisan and Gaspin, personal communication). Remarkably, each group matches a typical genomic context. The counterparts of P. furiosus PF1131 cas6 gene are all located 5′ to a I-A1 module, whereas the counterparts of the P. furiosus cas6-2 gene are always isolated in pyrococcal genomes. Finally, the third cas6 gene group, absent in P. furiosus, P. specie NA2 and P. abyssi, is located 5′ to the subtype III-A module in P. yayanosii, P. horikoshii and P. species ST04. Altogether, two cas6 groups appear to be associated with either CRISPR/cas system subtype I-A or subtype III-A, and the third group is remote from any CRISPR/cas system.
As mentioned earlier, one cas6 and one cas4 genes appear in contexts other than CRISPR/cas systems in the six pyrococcal genomes. Because no functional connections with CRISPR/cas systems are observed, caution should be used in classification as noted by Makarova et al.Citation7 These distant Cas4 and Cas6 might be involved in other uncharacterized cellular functions.
Among the six sequenced pyrococcal genomes, two distinct groups of CRISPR arrays can be distinguished on the basis of length and sequence of their respective direct repeats (DRs, and ). Note that the CRISPR arrays are extracted from CRISPRdbCitation1 and that some are reoriented to maintain most conserved repeats proximal to the leader sequence. Group 1 and Group 2 DRs are respectively 30 and 29 nucleotide long and have slightly differing consensus sequences (). CRISPR arrays tend to contain more Group 1 DRs than Group 2 DRs, the latter being absent from P. species NA2 and P. furiosus genomes (). CRISPR arrays with Group 1 DRs are associated to subtype I-A and III-B clusters, whereas CRISPR arrays adjacent to subtype III-A clusters contain Group 2 DRs (). Interestingly, the Group 2 DR () is related, by means of its consensus sequence and size, to one of the direct repeat groups reported as the “Unfolded Archaeal Cluster 6,”Citation52 which was assigned to the Mtube or subtype III-ACitation7,Citation49 in an analysis of CRISPR repeats of 195 microbial genomes. In general, the leader proximal repeats are more conserved than those situated at the end of an array.Citation53 As detailed below, the last eight DR nucleotides are due to become the 5′ tag of mature crRNAs. Interestingly, each DR group harbors a specific 5′ tag sequence that might direct the corresponding crRNAs to a different type of effector complex (). The spacer number per CRISPR array varies from 4–85 in pyrococcal genomes and their respective length ranges from 26–59 nucleotides. Because new spacers are acquired by active CRISPR/cas systems, these sequences may provide a record of the past “infection history.” A recent search for similarities to archaeal guide sequences in public databases, which reports potential signatures of genetic invaders in archaeal genomes,Citation54 did not recover any matching hit within pyrococcal spacers. With the exception of two short regions (14 and 17 bp) of two P. abyssi CRISPR 1 guides that are complementary to two coding sequences of PAV1,Citation55 no other similarity to thermococcal viruses or plasmids has been identified so far. Thus, the origin of pyrococcal CRISPR guide sequences remains to be determined.
Table 1. Two groups of direct repeats in pyrococcal CRISPR arrays.
The CRISPR arrays are preceded by an adenine/thymine (AT)-rich leader sequenceCitation56 that has been shown to contain promoter elementsCitation53,Citation57 and binding sites for regulatory proteins.Citation58 Leader sequences are also expected to direct the insertion of new guide sequences at the first DR during the adaptation phase.Citation53 Sequence alignments of the leaders preceding the 32 pyrococcal CRISPR arrays allow their clustering into two families (), which correspond to the two DR groups (). The BRE/TATA-box promoter signal is clearly defined for the Group 1 CRISPR arrays whereas only a TATA box signal, situated further away from the first DR, can be identified in leaders of Group 2 arrays. Group 2 leaders also have a highly conserved region 80 nucleotides upstream the TATA box that may correspond to a regulatory site ().
Figure 3. Putative promoter sequences found in pyrococcal leader regions. Manual alignment of the regions upstream the first repeat of each CRISPR array is shown. Two groups, corresponding to the two DR groups (see ) are delineated upon sequence conservation. The first repeats are shown in blue. Strictly conserved sequences are shown in red. Putative transcriptional signals (BRE/TATA elements) are indicated. The red arrows indicate the 5′ transcriptional starts experimentally identified in P. furiousCitation57 (above) and in P. abyssiCitation55 (below).

CRISPR Arrays Expression
The production of mature crRNAs that contain the invader targeting sequence is the second stage of CRISPR/cas-mediated immunity. Most CRISPR arrays are transcribed from their leader into long pre-crRNAs (pre-CRISPR RNAs), which are subsequently cleaved at each repeat to generate individual crRNAs. However, CRISPR systems are silent in a number of species. For instance, subtype I-E CRISPR systems are cryptic in most E. coli strains through a strong H-NS-mediated repression exerted on both the leaders and the promoter of the operon coding for the Cascade (subtype I-E effector) complex.Citation58 CRISPR array expression has been investigated in two different Pyrococcales: P. furiosus and P. abyssi. Multiple small RNA species (crRNA of various sizes) are expressed from all seven CRISPR loci in P. furiosus.Citation57,Citation59 In contrast, transcripts could only be detected from two of the four P. abyssi CRISPR arrays, indicating that not all CRISPR arrays are active in this species.Citation55 Only P. abyssi CRISPR 1 and 4, belonging to Group1 CRISPR arrays, were shown to be expressed from exponential through stationary growth phases. Transcription start sites have been mapped 24 nucleotides and 17 nucleotides downstream the TATA box in P. furiosus and P. abyssi, respectively ().Citation55,Citation57
Processing of Pre-crRNA by Cas6 Endoribonuclease
RNA analysis in P. furiosus and P. abyssi provided evidences that the pre-crRNA transcripts are indeed cleaved within the repeat sequences to release individual crRNA intermediates.Citation51,Citation55 The pre-crRNA processing differs in the three CRISPR types. While Type I and III systems use the Cas6 endoribonuclease,Citation50 either embedded in the effector complex (such as E. coli subtype I-E Cascade complexCitation60,Citation61) or alone (as detailed below for P. furiosus), pre-crRNAs are processed in Type II systems by the concerted action of Cas9, a tracrRNA (specific RNA complementary to the repeat sequence) and the cellular RNase III.Citation62
P. furiosus (Pf)Cas6 activity was shown to be metal-independent and to have a predicted active site similar to that of tRNA splicing endonucleasesCitation50,Citation63,Citation64 that generates 5′ hydroxyl and 2’-3′ cyclic phosphate ends.Citation50 RNA mutational analyses showed that PfCas6 binds to a 7-nucleotide sequence near the 5′ end of the repeat and cleaves 14 nucleotides downstream. The cleavage occurs between adenosine 22 and adenosine 23 of the 30-nucleotide repeat in P. furiosus and P. abyssi.Citation50,Citation55 Substitution of these two adenosines in the P. furiosus repeat disrupts cleavage but not PfCas6 binding.Citation50 A potential secondary structure of P. furiosus repeat was initially proposed,Citation50 but further RNA footprinting experiments revealed that the RNA repeat was unstructured in solution.Citation63 Furthermore, in addition to its binding site on the 5′ end of the repeat, PfCas6 also interacts with part of the 3′ end of the DR. Thus PfCas6 binds unstructured pre-crRNA in a wrap-around mechanism.Citation50,Citation63,Citation64
The crystal structure of PfCas6 revealed duplicated ferredoxin folds separated by a central cleft that contains a conserved glycine-rich loop and that the trans-esterification reaction operates through a general acid-base chemistry.Citation63 This has also been reported for Cas6 functional analogs from other CRISPR types such as E. coli Type I-E Cas6e (Cse3 or CasE)Citation60 or P. aeruginosa subtype I-F Cas6f (Csy4),Citation61 which share little homology with PfCas6 but all exhibit the ferredoxin fold or RNA recognition motif (RRM).Citation65 However, in contrast to PfCas6, Cas6e and Cas6f have been shown to recognize a stable hairpin structure formed by a pseudo-palindromic sequence upstream the cleavage site.Citation60,Citation61 Whichever the CRISPR system type (I or III), the resulting crRNAs all retain eight nucleotides of the repeat upstream of the guide sequence with no similarity between primary sequences of the CRISPR repeat.Citation51,Citation60,Citation61,Citation66 These eight nucleotides constitute the 5′ tag of crRNAs. The E. coli Cas6e operates as a subunit of a larger complex (Cascade) that binds to the mature crRNA to form the effector RNP complex implicated in the interference phase. On the other hand, PfCas6 acts aloneCitation63 and releases the resulting crRNAs, which are subsequently incorporated into an effector complex such as Cmr as detailed below.Citation51 P. furiosus guide sequences (inserts between two repeats) range from 34–59 nucleotides and Cas6-processed crRNAs range from 30–70 nucleotides in length. In P. furiosus, the intermediate crRNAs are further processed to abundant stable 35–45 nucleotide mature crRNAs.Citation59 In P. abyssi, discrete crRNA species in the same length range are also produced.Citation55 Cas6 establishes the 5′ end of mature crRNAs whereas the 3′ end is processed further by an unknown process.
The CMR Complex from P. furiosus Targets RNA
Large ribonucleoprotein (RNP) complexes could be isolated from P. furiosus cellular extracts on the basis of crRNA fractionation profiles.Citation51 Mass spectrometry of the purified complexes identified seven Cmr proteins [Cmr1-1, Cmr1-2 (two cmr1 genes), Cmr2 (Cas10), Cmr3, Cmr4, Cmr5 and Cmr6], all encoded by the RAMP (repeat-associated mysterious proteins) module,Citation49 which was renamed CRISPR/cas system subtype III-B,Citation7 adjacent to the I-A1 module and CRISPR locus 7 (). The crRNAs associated with these complexes and originated from all seven P. furiosus CRISPR loci present two types of mature crRNA species of 45 and 39 nucleotides with 5′ OH and 3′ phosphate end groups.Citation57 Overall, both crRNA sizes are represented in equivalent proportions in Cmr RNP complexes but individual guide RNA species can present altered ratios. The Cmr crRNA from P. furiosus all have in common the conserved 5′ tag sequence that results from Cas6 cleavageCitation50 and contains the last eight nucleotides from the repeat sequence (). The crRNAs two possible sizes (39 and 45 nucleotides) originate from differential trimming of their 3′ ends. The origin of this 3′ end trimming remains unknown. Evidence suggests that the sizes of Cmr crRNAs are defined by the distance from the 5′ end, independently of their sequences.Citation57 Thus, 3′ end trimming post-Cas6-cleavage could either be performed by a yet-to-be-determined process prior to incorporation into Cmr complexes, which would select the 39 and 45 nucleotides-long crRNA among the various possible sizes; or this size reduction could be performed by the Cmr complex itself upon binding to full-length crRNAs.
To date, the structural organization of the Cmr complex and the function of its individual components remain to be determined. The multidomain P. furiosus Cmr2 protein, signature protein in Type III CRISPR systems belonging to the Cas10 family, is the largest of the six proteins of the Cmr complex and was thought to be the catalytic subunit of the effector complex because of its N-terminal permutated histidine-aspartate superfamily hydrolase domain (HD) until it was recently shown that this domain is not essential for cleavage.Citation67 This suggested that another component of the Cmr complex provided the catalytic function.Citation67 The crystal structure of Cmr2 lacking the HD domain revealed other structural domains such as zinc finger domain, ferredoxin-like fold (polymerase/cyclase) nucleotide cyclase domain and two helical thumb domain of A-family DNA polymerase enzyme,Citation67,Citation68 but no specific activity could be identified for this Cmr component.
Nevertheless, the activity of the complex has been precisely characterized. Contrary to activities characterized for other Cas/crRNA complexes from Type I,Citation60 Type IICitation69 and subtype III-A,Citation66 which act on DNA substrates, no detectable effect on single-stranded or dsDNA substrates could be observed.Citation51 However, Cmr RNP complexes cleave ssRNA that are complementary to the crRNA held in the complex. This RNA targeting has been confirmed in vitro, using complexes purified from cell extracts or reconstituted from recombinant proteins.Citation51 Cmr complex activity against RNA was also confirmed in vivo using a fortuitous promoter in the second guide sequence from CRISPR locus 1 that transcribes backward toward the leader. The produced transcript is thus a substrate for Cmr RNP complexes using crRNA from the first guide. Northern blot analyses confirmed the transcript was cleaved in vivo.Citation57
Reconstitution of Cmr RNP complexes using recombinant proteins indicated that a single crRNA associated to Cmr1-1, Cmr2, Cmr3, Cmr4 and Cmr6 was sufficient to produce cleavage of a target RNA.Citation51 Thus, Cmr1-1 and Cmr1-2 are redundant and probably associate interchangeably to the Cmr complex. Cmr5 is also not necessary for target cleavage and its function remains to be determined. The RNA product can remain associated to some extent with the Cmr complex following cleavage.Citation57
At least 14 nucleotides of base pairing between the 3′ end of the crRNA and the target RNA molecule are necessary for recognition and cleavage of the target.Citation51 The Cmr ribonuclease effector complexes generate products with 3′ phosphate (or 2’-3′ cyclic phosphate) and 5′ hydroxyl termini, and require one of Mg2+, Mn2+, Ca2+, Zn2+, Ni2+ or Fe2+ divalent metal ions for activity. The complementary target RNAs are cleaved at a fixed 14 nucleotides distance from the guide crRNA 3′ end. This results in two possible cleavage sites of the target RNA depending on the guide crRNA size held by the Cmr complex. The presence of a functional 5′ tag on the crRNA is necessary for Cmr complex activity but it is not used to discriminate self from foreign RNA. The Cmr complex does not require a lack of complementarity between the target and the crRNA repeat tag, as seen with subtype III-A system.Citation66 Contrary to what is observed for type I and type II systems,Citation70 a specific PAM (protospacer adjacent motif) sequence does not seem to be required for Cmr complex target cleavage either. Thus, the Cmr RNP complexes do not discriminate self from non-self RNA targets.
Cmr systems are not universal in Pyrococcales. They are only found in P. furiosus and P. yayanosii genomes (). Overall, Cmr systems are found in 70% and 30% of archaeal and bacterial genomes, respectively.Citation71 A Cmr complex has been structurally and biochemically characterized from Sulfolobus solfataricus.Citation72 Although the Cmr systems from P. furiosus and S. solfataricus are homologous and both target RNA molecules complementary to their cognate crRNAs, they function differently. S. solfataricus Cmr complex is composed of 7 Cmr proteins and does not cleave its target at a fixed distance from its crRNA 3′ end, but at UA dinucleotides cleaving both target and crRNA strands.
Similarly to what has been proposed with other CRISPR systems,Citation73 Cmr complexes can easily be engineeredCitation57 to target any desired RNA sequence in a genetically tractable strain from the three domains of life, including eukaryotic cells. Subtype III-B systems thus have the potential to become a novel gene silencing tool with applications for industrial and medical research.
Discussion
Pyrococcales are a group of Thermococcales that are widely found in oceanic hyperthermophilic environments such as deep hydrothermal vents. There are a large number of reported thermococcal strains (more than 40) and possibly a large number of associated plasmids.Citation13 However, very few viruses have been reported so far for this group of archaea. Nevertheless, as found in 90% of archaeal genomes, all available pyrococcal genomes present at least one CRISPR/cas system. Similarly to what has been reported in the genomes of crenarchaeal Pyrobaculum speciesCitation48 and crenarchaeal Sulfolobales,Citation46 three different cas gene types were identified within the six pyrococcal genomes: a subtype I-A, a subtype III-A and a subtype III-B system (). Typically, subtype I-A is ubiquitously represented in Pyrococcales.
However, a closer look suggests that pyrococcal CRISPR systems fall into two categories, associated either with subtype I-A or III-A systems. Each group seems to have its own set of leader sequences and direct repeats (Group 1 and 2 respectively), informational module (cas1, cas2 and cas4 with dedicated clustering), crRNA processing Cas6 and effector Cas complexes that are separated into multiple modules (). Multiple effector clusters are found associated to subtype I-A systems, including the I-A1 and I-A2 modules. These gene clusters seem to be two homologous subtype I-A effector modules, characterized by the presence of either cas8a1/cst1 or cas8a2. We suggest that subtype III-B genes (a.k.a. RAMP or cmr genes) might be viewed as a subtype I-A effector module, rather than an autonomous CRISPR/cas system. For instance, in the case of pyrococcal genomes, no informational module or CRISPR array are ever found associated to subtype III-B gene clusters, except when they form a subtype I-A/III-B hybrid system where they are associated to the subtype I-A informational module and CRISPR array (as found in P. furiosus).
Figure 4. Model of pyrococcal CRISPR mode of action. Subtype I-A and III-B systems are delineated with their respective CRISPR arrays, cas6 (cyan) and informational modules (cas1, cas2 and cas4 in blue). The various guide sequences are indicated in green. Group 1 direct repeats (DRs), associated to the subtype I-A system, are represented in orange, whereas the Group 2 DRs, connected to the subtype III-A system are in pink. Effector cas genes and their products, which associate to crRNAs to form the interference complexes, are represented in yellow for subtype I-A, red for subtype III-B and purple for subtype III-A. Subtype I-A and III-B RNP complexes are hypothesized to associate to the same population of crRNAs (from Group 1 CRISPR arrays). Type III-B RNP complexes carry exclusively 39- and 45-nucleotide long crRNA and are the only CRISPR RNP complexes known to target RNA molecules.Citation57 As processed crRNA of other sizes are also found in vivo, they are expected to be associated with the subtype I-A RNP complexes, although association of the 39- and 45-nucleotide long crRNA to this complex is not excluded. On the other hand, subtype III-A RNP complexes are expected to be associated with crRNAs from the Group 2 CRISPR arrays. Further experiments remain to be performed to validate this model.
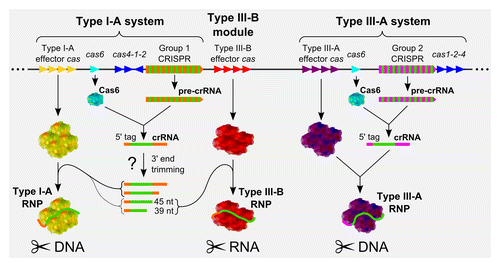
As opposed to all other CRISPR systems that target DNA, subtype III-B effector RNP complex is the only one known to target RNA molecules. This subtype III-B module could be viewed as an option, for the subtype I-A CRISPR system, conferring the ability to target RNA in addition to DNA, using the same set of guide sequences (). However, the fact that I-A and III-B subtype effector RNP complexes use the same set of crRNAs to target either DNA or RNA remains to be confirmed experimentally.
The existence of such an RNA-targeting system is intriguing as no archaeal RNA virus has been identified so far. This raises the question: why has such an RNA targeting module developed? Is there a benefit for a hyperthermophilic archaeal cell to degrade RNA molecules from an invader? There might be some RNA viruses from deep vents that we have not yet identified. This system could alter the expression of infecting genetic elements by silencing their mRNAs while other defense systems (CRISPR or not) rid the cell from the invader or trigger programmed cell death, thus preventing the infectious genetic element from spreading across the population. This subtype III-B system could also have functions other than anti-viral defense, such as gene regulation, although guides targeting cellular genes are the exception rather than the rule.
Although no experimental evidence exists so far, subtype I-A and subtype III-B are hypothesized to use the same set of crRNAs (from group 1 CRISPR arrays), whereas Type III-A complexes seem to be independent and to use their own set of crRNAs (from group 2 CRISPR arrays, which are absent from P. furiosus and P. species NA2 genomes). Group 1 and group 2 crRNAs contain different 5′ tags () that hypothetically allow each effector complex to select its dedicated crRNA after their processing by Group 1 or 2 specific Cas6 endoribonucleases (). This situation could be experimentally tested in P. yayanosii, which contains the whole set of CRISPR modules (I-A1, I-A2, III-B and III-A) and the two groups of CRISPR arrays ().
The P. abyssi genome differs from the other Pyrococcales. It is intriguing because it does not present a single full CRISPR system (). The P. abyssi genome contains the I-A2 effector module but lacks the associated informational module and more specifically cas1 and cas2 genes, which theoretically impairs this species from acquiring new guide sequences. P. abyssi carries a group 2 CRISPR array, indicating it might have once possessed a functional subtype III-A system, which could have been lost afterward. Thus, the intriguing P. abyssi CRISPR profile is most probably the result of gene loss. This organism possibly once had a more elaborated CRISPR system organization and might have lost several of its CRISPR components. This raises the question of whether P. abyssi has CRISPR activity. Experimental evidences show that two of the four CRISPR arrays are expressed (CRISPR 1 and 4) and their transcripts processed into individual crRNAs. These two arrays are of the subtype I-A group for which cas genes (I-A2 module) are still present in the strain (). Thus, it is probable that this residual CRISPR system can still target invaders bearing some complementarity to the set of guides from arrays 1 and 4 but cannot develop immunity toward novel genetic invaders, since cas1 and cas2 appear to miss. It would be interesting to test this hypothesis. A functional adaptation module could be restored through genetic manipulation of P. abyssi genome, which is now feasible. Both wild-type and this modified strain could be tested for their ability to acquire new guides. If it appeared to be the case, the possibility that P. abyssi would uptake novel spacers, within its CRISPR arrays, without the need for Cas1 and Cas2 proteins would be truly interesting as this would open new opportunities to understand how guide sequences are selected and integrated in CRISPR arrays.
The origins of pyrococcal CRISPR guide sequences remain to be determined. The lack of matching hit results from the scarcity of available thermococcal plasmid and virus sequences. The ever increasing number of available sequences will at some point allow the discovery of other matches between thermococcal CRISPR guides and genetic invaders, providing additional evidence that CRISPR systems participate in the defense against plasmids and viruses.
The diversity of CRISPR system situations in the Pyrococcales illustrates the modularity and instability of CRISPR systems. It would be quite challenging to decipher the ancestral CRISPR organization in the Pyrococcales. One possibility is that the common ancestor to all Pyrococcales (and possibly Thermococcales) had all three CRISPR subtype systems, which would have then been differentially lost over time, depending on the species (e.g., P. abyssi or P. species NA2). The opposite possibility is that the common ancestor had no CRISPR at all and species have been differentially “infected” by various CRISPR systems that could have been brought to them by mobile genetic elements, such as plasmids, which have been reported to also carry CRISPR systems.Citation74,Citation75
Thus CRISPR systems in the Pyrococcales are diverse and dynamic. There are a lot of things that we still do not understand on the pyrococcal CRISPR/cas systems organization and, since the lack of experimental data, even less on Thermococcus systems. But knowledge in the CRISPR field is added as quickly as guide sequences in an active CRISPR array. Hopefully, future experimental work on Thermococcus CRISPR/cas systems will allow accurate cas gene annotations and extend our global knowledge of CRISPR/cas systems of the Thermococcal order. And the better we understand these CRISPR systems, the more we can exploit them, either as tools for molecular biology, or as ways to fight viral infections in prokaryotic populations of interest to industries.
| Abbreviations: | ||
| CRISPR | = | clustered regularly interspaced short palindromic repeats |
| bp | = | base pair |
| kb | = | kilo base |
| ds | = | double strand |
| ss | = | single strand |
| cas | = | CRISPR associated |
| crRNA | = | CRISPR RNA |
| RNP | = | ribonucleoprotein |
Acknowledgments
We are grateful to Y. Quentin for the thermococcal phylogenetic tree presented in . We thank A.J. Carpousis for critical reading of the manuscript. This work was supported by the Centre National de la Recherche Scientifique (CNRS) with additional funding from the Agence Nationale de la Recherche (ANR) (BLAN08-1_329396) (B.C.-O.). We thank the GenoToul Bioinformatics Platform for providing databanks access and computing resources.
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
References
- Grissa I, Vergnaud G, Pourcel C. The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats. BMC Bioinformatics 2007; 8:172; http://dx.doi.org/10.1186/1471-2105-8-172; PMID: 17521438
- Karginov FV, Hannon GJ. The CRISPR system: small RNA-guided defense in bacteria and archaea. Mol Cell 2010; 37:7 - 19; http://dx.doi.org/10.1016/j.molcel.2009.12.033; PMID: 20129051
- Bhaya D, Davison M, Barrangou R. CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu Rev Genet 2011; 45:273 - 97; http://dx.doi.org/10.1146/annurev-genet-110410-132430; PMID: 22060043
- Westra ER, Swarts DC, Staals RH, Jore MM, Brouns SJ, van der Oost J. The CRISPRs, they are a-changin’: how prokaryotes generate adaptive immunity. Annu Rev Genet 2012; 46:311 - 39; http://dx.doi.org/10.1146/annurev-genet-110711-155447; PMID: 23145983
- Wiedenheft B, Sternberg SH, Doudna JA. RNA-guided genetic silencing systems in bacteria and archaea. Nature 2012; 482:331 - 8; http://dx.doi.org/10.1038/nature10886; PMID: 22337052
- Makarova KS, Aravind L, Wolf YI, Koonin EV. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol Direct 2011; 6:38; http://dx.doi.org/10.1186/1745-6150-6-38; PMID: 21756346
- Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P, et al. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 2011; 9:467 - 77; http://dx.doi.org/10.1038/nrmicro2577; PMID: 21552286
- Kecha M, Benallaoua S, Touzel JP, Bonaly R, Duchiron F. Biochemical and phylogenetic characterization of a novel terrestrial hyperthermophilic archaeon pertaining to the genus Pyrococcus from an Algerian hydrothermal hot spring. Extremophiles 2007; 11:65 - 73; http://dx.doi.org/10.1007/s00792-006-0010-9; PMID: 16969710
- Mardanov AV, Ravin NV, Svetlitchnyi VA, Beletsky AV, Miroshnichenko ML, Bonch-Osmolovskaya EA, et al. Metabolic versatility and indigenous origin of the archaeon Thermococcus sibiricus, isolated from a siberian oil reservoir, as revealed by genome analysis. Appl Environ Microbiol 2009; 75:4580 - 8; http://dx.doi.org/10.1128/AEM.00718-09; PMID: 19447963
- Miroshnichenko ML, Hippe H, Stackebrandt E, Kostrikina NA, Chernyh NA, Jeanthon C, et al. Isolation and characterization of Thermococcus sibiricus sp. nov. from a Western Siberia high-temperature oil reservoir. Extremophiles 2001; 5:85 - 91; http://dx.doi.org/10.1007/s007920100175; PMID: 11354459
- Roussel EG, Sauvadet AL, Chaduteau C, Fouquet Y, Charlou JL, Prieur D, et al. Archaeal communities associated with shallow to deep subseafloor sediments of the New Caledonia Basin. Environ Microbiol 2009; 11:2446 - 62; http://dx.doi.org/10.1111/j.1462-2920.2009.01976.x; PMID: 19624712
- Takai K, Gamo T, Tsunogai U, Nakayama N, Hirayama H, Nealson KH, et al. Geochemical and microbiological evidence for a hydrogen-based, hyperthermophilic subsurface lithoautotrophic microbial ecosystem (HyperSLiME) beneath an active deep-sea hydrothermal field. Extremophiles 2004; 8:269 - 82; http://dx.doi.org/10.1007/s00792-004-0386-3; PMID: 15309563
- Prieur D, Erauso G, Flament D, Gaillard M, Geslin C, Gonnet M, et al. Deep-sea Thermococcales and their genetic elements: plasmids and viruses. Methods in Microbiology 2006; 35:253 - 78; http://dx.doi.org/10.1016/S0580-9517(08)70014-X
- Cohen GN, Barbe V, Flament D, Galperin M, Heilig R, Lecompte O, et al. An integrated analysis of the genome of the hyperthermophilic archaeon Pyrococcus abyssi. Mol Microbiol 2003; 47:1495 - 512; http://dx.doi.org/10.1046/j.1365-2958.2003.03381.x; PMID: 12622808
- Kawarabayasi Y, Sawada M, Horikawa H, Haikawa Y, Hino Y, Yamamoto S, et al. Complete sequence and gene organization of the genome of a hyper-thermophilic archaebacterium, Pyrococcus horikoshii OT3. DNA Res 1998; 5:55 - 76; http://dx.doi.org/10.1093/dnares/5.2.55; PMID: 9679194
- Robb FT, Maeder DL, Brown JR, DiRuggiero J, Stump MD, Yeh RK, et al. Genomic sequence of hyperthermophile, Pyrococcus furiosus: implications for physiology and enzymology. Methods Enzymol 2001; 330:134 - 57; http://dx.doi.org/10.1016/S0076-6879(01)30372-5; PMID: 11210495
- Jun X, Lupeng L, Minjuan X, Oger P, Fengping W, Jebbar M, et al. Complete genome sequence of the obligate piezophilic hyperthermophilic archaeon Pyrococcus yayanosii CH1. J Bacteriol 2011; 193:4297 - 8; http://dx.doi.org/10.1128/JB.05345-11; PMID: 21705594
- Lee HS, Bae SS, Kim MS, Kwon KK, Kang SG, Lee JH. Complete genome sequence of hyperthermophilic Pyrococcus sp. strain NA2, isolated from a deep-sea hydrothermal vent area. J Bacteriol 2011; 193:3666 - 7; http://dx.doi.org/10.1128/JB.05150-11; PMID: 21602357
- Jung JH, Lee JH, Holden JF, Seo DH, Shin H, Kim HY, et al. Complete genome sequence of the hyperthermophilic archaeon Pyrococcus sp. strain ST04, isolated from a deep-sea hydrothermal sulfide chimney on the Juan de Fuca Ridge. J Bacteriol 2012; 194:4434 - 5; http://dx.doi.org/10.1128/JB.00824-12; PMID: 22843576
- Vannier P, Marteinsson VT, Fridjonsson OH, Oger P, Jebbar M. Complete genome sequence of the hyperthermophilic, piezophilic, heterotrophic, and carboxydotrophic archaeon Thermococcus barophilus MP. J Bacteriol 2011; 193:1481 - 2; http://dx.doi.org/10.1128/JB.01490-10; PMID: 21217005
- Zivanovic Y, Armengaud J, Lagorce A, Leplat C, Guérin P, Dutertre M, et al. Genome analysis and genome-wide proteomics of Thermococcus gammatolerans, the most radioresistant organism known amongst the Archaea. Genome Biol 2009; 10:R70; http://dx.doi.org/10.1186/gb-2009-10-6-r70; PMID: 19558674
- Oger P, Sokolova TG, Kozhevnikova DA, Chernyh NA, Bartlett DH, Bonch-Osmolovskaya EA, et al. Complete genome sequence of the hyperthermophilic archaeon Thermococcus sp. strain AM4, capable of organotrophic growth and growth at the expense of hydrogenogenic or sulfidogenic oxidation of carbon monoxide. J Bacteriol 2011; 193:7019 - 20; http://dx.doi.org/10.1128/JB.06259-11; PMID: 22123768
- Wang X, Gao Z, Xu X, Ruan L. Complete genome sequence of Thermococcus sp. strain 4557, a hyperthermophilic archaeon isolated from a deep-sea hydrothermal vent area. J Bacteriol 2011; 193:5544 - 5; http://dx.doi.org/10.1128/JB.05851-11; PMID: 21914870
- Jung JH, Holden JF, Seo DH, Park KH, Shin H, Ryu S, et al. Complete genome sequence of the hyperthermophilic archaeon Thermococcus sp. strain CL1, isolated from a Paralvinella sp. polychaete worm collected from a hydrothermal vent. J Bacteriol 2012; 194:4769 - 70; http://dx.doi.org/10.1128/JB.01016-12; PMID: 22887670
- Zivanovic Y, Lopez P, Philippe H, Forterre P. Pyrococcus genome comparison evidences chromosome shuffling-driven evolution. Nucleic Acids Res 2002; 30:1902 - 10; http://dx.doi.org/10.1093/nar/30.9.1902; PMID: 11972326
- Pina M, Bize A, Forterre P, Prangishvili D. The archeoviruses. FEMS Microbiol Rev 2011; 35:1035 - 54; http://dx.doi.org/10.1111/j.1574-6976.2011.00280.x; PMID: 21569059
- Peng X, Garrett RA, She Q. Archaeal viruses–novel, diverse and enigmatic. Science China. Life Sci 2012; 55:422 - 33; http://dx.doi.org/10.1007/s11427-012-4325-8
- Pietilä MK, Roine E, Paulin L, Kalkkinen N, Bamford DH. An ssDNA virus infecting archaea: a new lineage of viruses with a membrane envelope. Mol Microbiol 2009; 72:307 - 19; http://dx.doi.org/10.1111/j.1365-2958.2009.06642.x; PMID: 19298373
- Mochizuki T, Krupovic M, Pehau-Arnaudet G, Sako Y, Forterre P, Prangishvili D. Archaeal virus with exceptional virion architecture and the largest single-stranded DNA genome. Proc Natl Acad Sci USA 2012; 109:13386 - 91; http://dx.doi.org/10.1073/pnas.1203668109; PMID: 22826255
- Benbouzid-Rollet N, López-García P, Watrin L, Erauso G, Prieur D, Forterre P. Isolation of new plasmids from hyperthermophilic Archaea of the order Thermococcales. Res Microbiol 1997; 148:767 - 75; http://dx.doi.org/10.1016/S0923-2508(97)82452-7; PMID: 9765860
- Lepage E, Marguet E, Geslin C, Matte-Tailliez O, Zillig W, Forterre P, et al. Molecular diversity of new Thermococcales isolates from a single area of hydrothermal deep-sea vents as revealed by randomly amplified polymorphic DNA fingerprinting and 16S rRNA gene sequence analysis. Appl Environ Microbiol 2004; 70:1277 - 86; http://dx.doi.org/10.1128/AEM.70.3.1277-1286.2004; PMID: 15006744
- Erauso G, Marsin S, Benbouzid-Rollet N, Baucher MF, Barbeyron T, Zivanovic Y, et al. Sequence of plasmid pGT5 from the archaeon Pyrococcus abyssi: evidence for rolling-circle replication in a hyperthermophile. J Bacteriol 1996; 178:3232 - 7; PMID: 8655503
- Gonnet M, Erauso G, Prieur D, Le Romancer M. pAMT11, a novel plasmid isolated from a Thermococcus sp. strain closely related to the virus-like integrated element TKV1 of the Thermococcus kodakaraensis genome. Res Microbiol 2011; 162:132 - 43; http://dx.doi.org/10.1016/j.resmic.2010.11.003; PMID: 21144896
- Ward DE, Revet IM, Nandakumar R, Tuttle JH, de Vos WM, van der Oost J, et al. Characterization of plasmid pRT1 from Pyrococcus sp. strain JT1. J Bacteriol 2002; 184:2561 - 6; http://dx.doi.org/10.1128/JB.184.9.2561-2566.2002; PMID: 11948174
- Soler N, Justome A, Quevillon-Cheruel S, Lorieux F, Le Cam E, Marguet E, et al. The rolling-circle plasmid pTN1 from the hyperthermophilic archaeon Thermococcus nautilus. Mol Microbiol 2007; 66:357 - 70; http://dx.doi.org/10.1111/j.1365-2958.2007.05912.x; PMID: 17784911
- Soler N, Marguet E, Cortez D, Desnoues N, Keller J, van Tilbeurgh H, et al. Two novel families of plasmids from hyperthermophilic archaea encoding new families of replication proteins. Nucleic Acids Res 2010; 38:5088 - 104; http://dx.doi.org/10.1093/nar/gkq236; PMID: 20403814
- Krupovic M, Gonnet M, Hania WB, Forterre P, Erauso G. Insights into dynamics of mobile genetic elements in hyperthermophilic environments from five new thermococcus plasmids. PLoS One 2013; 8:e49044; http://dx.doi.org/10.1371/journal.pone.0049044; PMID: 23326305
- Geslin C, Le Romancer M, Erauso G, Gaillard M, Perrot G, Prieur D. PAV1, the first virus-like particle isolated from a hyperthermophilic euryarchaeote, “Pyrococcus abyssi”. J Bacteriol 2003; 185:3888 - 94; http://dx.doi.org/10.1128/JB.185.13.3888-3894.2003; PMID: 12813083
- Geslin C, Gaillard M, Flament D, Rouault K, Le Romancer M, Prieur D, et al. Analysis of the first genome of a hyperthermophilic marine virus-like particle, PAV1, isolated from Pyrococcus abyssi. J Bacteriol 2007; 189:4510 - 9; http://dx.doi.org/10.1128/JB.01896-06; PMID: 17449623
- Gorlas A, Koonin EV, Bienvenu N, Prieur D, Geslin C. TPV1, the first virus isolated from the hyperthermophilic genus Thermococcus. Environ Microbiol 2012; 14:503 - 16; http://dx.doi.org/10.1111/j.1462-2920.2011.02662.x; PMID: 22151304
- Fukui T, Atomi H, Kanai T, Matsumi R, Fujiwara S, Imanaka T. Complete genome sequence of the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1 and comparison with Pyrococcus genomes. Genome Res 2005; 15:352 - 63; http://dx.doi.org/10.1101/gr.3003105; PMID: 15710748
- Clore AJ, Stedman KM. The SSV1 viral integrase is not essential. Virology 2007; 361:103 - 11; http://dx.doi.org/10.1016/j.virol.2006.11.003; PMID: 17175004
- Benson DA, Karsch-Mizrachi I, Clark K, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res 2012; 40:Database issue D48 - 53; http://dx.doi.org/10.1093/nar/gkr1202; PMID: 22144687
- Chan PP, Holmes AD, Smith AM, Tran D, Lowe TM. The UCSC Archaeal Genome Browser: 2012 update. Nucleic Acids Res 2012; 40:Database issue D646 - 52; http://dx.doi.org/10.1093/nar/gkr990; PMID: 22080555
- Selengut JD, Haft DH, Davidsen T, Ganapathy A, Gwinn-Giglio M, Nelson WC, et al. TIGRFAMs and Genome Properties: tools for the assignment of molecular function and biological process in prokaryotic genomes. Nucleic Acids Res 2007; 35:Database issue D260 - 4; http://dx.doi.org/10.1093/nar/gkl1043; PMID: 17151080
- Garrett RA, Shah SA, Vestergaard G, Deng L, Gudbergsdottir S, Kenchappa CS, et al. CRISPR-based immune systems of the Sulfolobales: complexity and diversity. Biochem Soc Trans 2011; 39:51 - 7; http://dx.doi.org/10.1042/BST0390051; PMID: 21265746
- Garrett RA, Vestergaard G, Shah SA. Archaeal CRISPR-based immune systems: exchangeable functional modules. Trends Microbiol 2011; 19:549 - 56; http://dx.doi.org/10.1016/j.tim.2011.08.002; PMID: 21945420
- Bernick DL, Cox CL, Dennis PP, Lowe TM. Comparative genomic and transcriptional analyses of CRISPR systems across the genus Pyrobaculum. Front Microbiol 2012; 3:251; http://dx.doi.org/10.3389/fmicb.2012.00251; PMID: 22811677
- Haft DH, Selengut J, Mongodin EF, Nelson KE. A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput Biol 2005; 1:e60; http://dx.doi.org/10.1371/journal.pcbi.0010060; PMID: 16292354
- Carte J, Wang R, Li H, Terns RM, Terns MP. Cas6 is an endoribonuclease that generates guide RNAs for invader defense in prokaryotes. Genes Dev 2008; 22:3489 - 96; http://dx.doi.org/10.1101/gad.1742908; PMID: 19141480
- Hale CR, Zhao P, Olson S, Duff MO, Graveley BR, Wells L, et al. RNA-guided RNA cleavage by a CRISPR RNA-Cas protein complex. Cell 2009; 139:945 - 56; http://dx.doi.org/10.1016/j.cell.2009.07.040; PMID: 19945378
- Kunin V, Sorek R, Hugenholtz P. Evolutionary conservation of sequence and secondary structures in CRISPR repeats. Genome Biol 2007; 8:R61; http://dx.doi.org/10.1186/gb-2007-8-4-r61; PMID: 17442114
- Horvath P, Barrangou R. CRISPR/Cas, the immune system of bacteria and archaea. Science 2010; 327:167 - 70; http://dx.doi.org/10.1126/science.1179555; PMID: 20056882
- Brodt A, Lurie-Weinberger MN, Gophna U. CRISPR loci reveal networks of gene exchange in archaea. Biol Direct 2011; 6:65; http://dx.doi.org/10.1186/1745-6150-6-65; PMID: 22188759
- Phok K, Moisan A, Rinaldi D, Brucato N, Carpousis AJ, Gaspin C, et al. Identification of CRISPR and riboswitch related RNAs among novel noncoding RNAs of the euryarchaeon Pyrococcus abyssi. BMC Genomics 2011; 12:312; http://dx.doi.org/10.1186/1471-2164-12-312; PMID: 21668986
- Jansen R, Embden JD, Gaastra W, Schouls LM. Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 2002; 43:1565 - 75; http://dx.doi.org/10.1046/j.1365-2958.2002.02839.x; PMID: 11952905
- Hale CR, Majumdar S, Elmore J, Pfister N, Compton M, Olson S, et al. Essential features and rational design of CRISPR RNAs that function with the Cas RAMP module complex to cleave RNAs. Mol Cell 2012; 45:292 - 302; http://dx.doi.org/10.1016/j.molcel.2011.10.023; PMID: 22227116
- Pul U, Wurm R, Arslan Z, Geissen R, Hofmann N, Wagner R. Identification and characterization of E. coli CRISPR-cas promoters and their silencing by H-NS. Mol Microbiol 2010; 75:1495 - 512; http://dx.doi.org/10.1111/j.1365-2958.2010.07073.x; PMID: 20132443
- Hale C, Kleppe K, Terns RM, Terns MP. Prokaryotic silencing (psi)RNAs in Pyrococcus furiosus. RNA 2008; 14:2572 - 9; http://dx.doi.org/10.1261/rna.1246808; PMID: 18971321
- Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 2008; 321:960 - 4; http://dx.doi.org/10.1126/science.1159689; PMID: 18703739
- Haurwitz RE, Jinek M, Wiedenheft B, Zhou K, Doudna JA. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 2010; 329:1355 - 8; http://dx.doi.org/10.1126/science.1192272; PMID: 20829488
- Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 2011; 471:602 - 7; http://dx.doi.org/10.1038/nature09886; PMID: 21455174
- Carte J, Pfister NT, Compton MM, Terns RM, Terns MP. Binding and cleavage of CRISPR RNA by Cas6. RNA 2010; 16:2181 - 8; http://dx.doi.org/10.1261/rna.2230110; PMID: 20884784
- Wang R, Preamplume G, Terns MP, Terns RM, Li H. Interaction of the Cas6 riboendonuclease with CRISPR RNAs: recognition and cleavage. Structure 2011; 19:257 - 64; http://dx.doi.org/10.1016/j.str.2010.11.014; PMID: 21300293
- Maris C, Dominguez C, Allain FH. The RNA recognition motif, a plastic RNA-binding platform to regulate post-transcriptional gene expression. FEBS J 2005; 272:2118 - 31; http://dx.doi.org/10.1111/j.1742-4658.2005.04653.x; PMID: 15853797
- Marraffini LA, Sontheimer EJ. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 2008; 322:1843 - 5; http://dx.doi.org/10.1126/science.1165771; PMID: 19095942
- Cocozaki AI, Ramia NF, Shao Y, Hale CR, Terns RM, Terns MP, et al. Structure of the Cmr2 subunit of the CRISPR-Cas RNA silencing complex. Structure 2012; 20:545 - 53; http://dx.doi.org/10.1016/j.str.2012.01.018; PMID: 22405013
- Zhu X, Ye K. Crystal structure of Cmr2 suggests a nucleotide cyclase-related enzyme in type III CRISPR-Cas systems. FEBS Lett 2012; 586:939 - 45; http://dx.doi.org/10.1016/j.febslet.2012.02.036; PMID: 22449983
- Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P, et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 2010; 468:67 - 71; http://dx.doi.org/10.1038/nature09523; PMID: 21048762
- Mojica FJ, Díez-Villaseñor C, García-Martínez J, Almendros C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 2009; 155:733 - 40; http://dx.doi.org/10.1099/mic.0.023960-0; PMID: 19246744
- Shah SA, Garrett RA. CRISPR/Cas and Cmr modules, mobility and evolution of adaptive immune systems. Res Microbiol 2011; 162:27 - 38; http://dx.doi.org/10.1016/j.resmic.2010.09.001; PMID: 20863886
- Zhang J, Rouillon C, Kerou M, Reeks J, Brugger K, Graham S, et al. Structure and mechanism of the CMR complex for CRISPR-mediated antiviral immunity. Mol Cell 2012; 45:303 - 13; http://dx.doi.org/10.1016/j.molcel.2011.12.013; PMID: 22227115
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012; 337:816 - 21; http://dx.doi.org/10.1126/science.1225829; PMID: 22745249
- Guo P, Cheng Q, Xie P, Fan Y, Jiang W, Qin Z. Characterization of the multiple CRISPR loci on Streptomyces linear plasmid pSHK1. Acta Biochim Biophys Sin (Shanghai) 2011; 43:630 - 9; http://dx.doi.org/10.1093/abbs/gmr052; PMID: 21705768
- Yang Y, Kurokawa T, Takahama Y, Nindita Y, Mochizuki S, Arakawa K, et al. pSLA2-M of Streptomyces rochei is a composite linear plasmid characterized by self-defense genes and homology with pSLA2-L. Biosci Biotechnol Biochem 2011; 75:1147 - 53; http://dx.doi.org/10.1271/bbb.110054; PMID: 21670526
- Phung DK, Rinaldi D, Langendijk-Genevaux PS, Quentin Y, Carpousis AJ, Clouet-d’Orval B. Archaeal β-CASP ribonucleases of the aCPSF1 family are orthologs of the eukaryal CPSF-73 factor. Nucleic Acids Res 2013; 41:1091 - 103; http://dx.doi.org/10.1093/nar/gks1237; PMID: 23222134
- Crooks GE, Hon G, Chandonia JM, Brenner SE. WebLogo: a sequence logo generator. Genome Res 2004; 14:1188 - 90; http://dx.doi.org/10.1101/gr.849004; PMID: 15173120