Figures & data
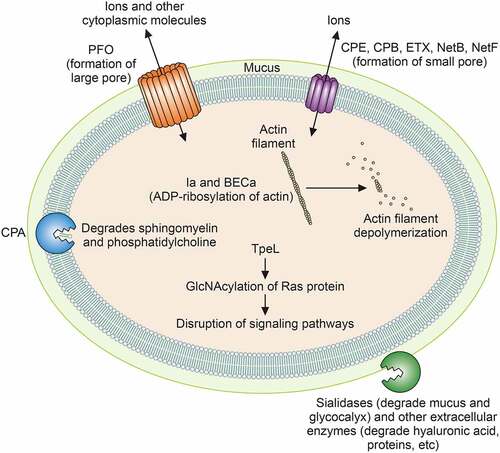
Table 1. Characteristics of toxins and extracellular degradative enzymes produced by C. perfringens
Table 2. Current C. perfringens toxinotyping scheme
Table 3. C. perfringens toxinotype: disease associations
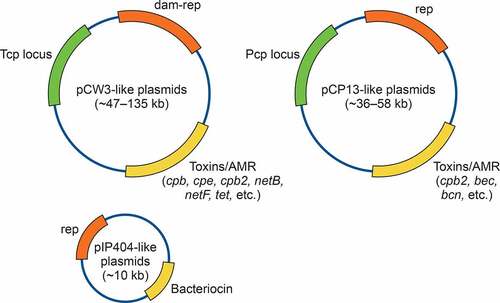
Table 4. Size and diversity C. perfringens plasmids encoding key-toxins1.
![Figure 3. Current model for cross-talk between the VirS/VirR two-component regulatory system and the Agr-like quorum sensing system in C. perfringens. The AgrD peptide is processed by AgrB (and perhaps other unidentified factors) to form a cyclic autoinducing signaling peptide (AIP). AIP then binds to VirS, which in turn phosphorylates (p) VirR. The phosphorylated VirR protein then binds to VirR boxes upstream of some toxins genes (e.g. the pfoA gene) and upstream of the vrr gene encoding VR-RNA. VR-RNA then leads to increased transcription of genes encoding toxins such as CPA. This reults in increased production of those toxins. Based upon [Citation162,Citation308]](/cms/asset/de7a3e1c-d36b-4a80-8d32-084c8a4315ff/kvir_a_1886777_f0003_oc.jpg)