Figures & data
Table 1. Bacterial strain characteristics and corresponding the MALDITet(X)-plus test results
Table 2. Characteristics of test strains used for test validation
Figure 1. Strategy for identification of Tet(x)-producers and non-Tet(X)-producing tetracycline-resistant strains and tetracycline-susceptible strains using the MALDITet(X)-plus test. Strains that showed peaks of tigecycline metabolite or oxytetracycline metabolite were labeled with “+”
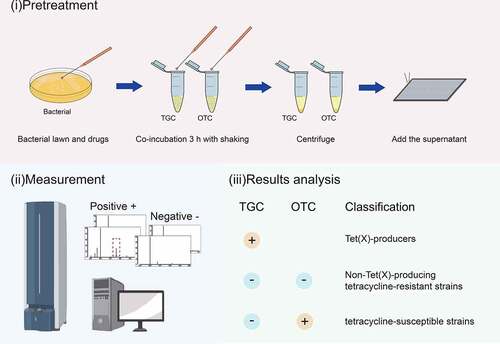
Figure 2. Representative MALDI-TOF MS results for detection of Tet(x)-producers and non-Tet(X)-producers. (a) Structures of TGC and OTC along with their oxygen-modified derivatives. (b) Representative MALDI-TOF MS spectra of tigecycline and oxytetracycline oxygenation assays after a 3 h incubation. Peaks of tigecycline and tigecycline metabolite are denoted by dashed red lines and represent peaks at m/z 586 ± 0.2 and m/z 602 ± 0.2, respectively. Peaks of oxytetracycline and oxytetracycline metabolite are denoted by dashed blue lines and represent peaks at m/z 460 ± 0.2 and m/z 476 ± 0.2, respectively
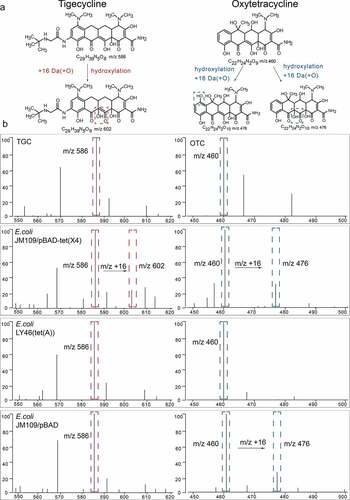
Figure 3. MALDITet(x)-plus test results using 319 test strains. (a) The cutoff value of 0.00405 can clearly distinguish Tet(X)-producers M/(M + T) >0.00405 (b) The cutoff value of 0.05945 can clearly identify non-Tet(X)-producers possessing M/(M + O) <0.05945. Three independent experiments were performed
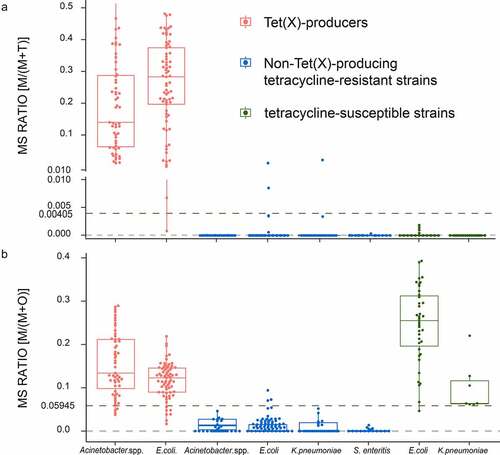
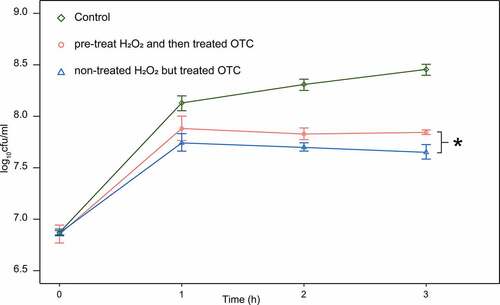
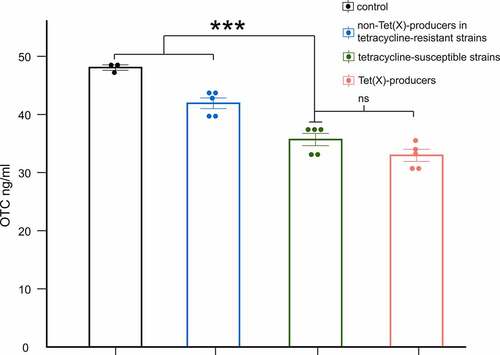
Figure 4. LC-MS/MS results for degradation of OTC by Tet(x)-producers, non-Tet(X) producers in tetracycline-resistant strains, and tetracycline-susceptible strains. The control group includes only OTC. We compare tetracycline-susceptible strains with Tet(X)-producers, non-Tet(X) producers in tetracycline-resistant strains, and the control group (three asterisk for p < 0.001)
Supplemental Material
Download Zip (359 KB)Data availability statement
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials.