ABSTRACT
This study aimed to elucidate the underlying mechanisms for the protective effect of Lactobacillus on hypercholesterolemia. Firstly, twenty Lactobacillus strains were screened for potential probiotics with high bile salt hydrolase (BSH) activity using in vitro methods. Secondly, hypercholesterolemic mice induced by a cholesterol-enriched diet were administrated by the probiotic candidate strain L. plantarum Y15, identified from the screen. Results in vitro showed that L. plantarum Y15 possessed high BSH activity. In vivo, L. plantarum Y15 supplementation decreased total cholesterol and triglyceride in serum and liver, increased total cholesterol and total bile acid in feces, and alleviated histopathological changes in the liver. Furthermore, L. plantarum Y15 supplementation modulated the gut microbiota to the control group’s pattern, leading to increase BSH activity, which further influence FXR and SHP signaling pathway to upregulate the expression level of CYP7A1 to enhance the cholesterol catabolism.
RESUMEN
Este estudio se propuso dilucidar los mecanismos subyacentes al efecto protector de Lactobacillus sobre la hipercolesterolemia. En primer lugar, se utilizaron métodos in vitro para examinar veinte cepas de Lactobacillus en busca de probióticos potenciales con alta actividad de hidrolasa de sales biliares (BSH). Luego se administró la cepa candidata a probiótico L. plantarum Y15, identificada a partir del cribado, a ratones hipercolesterolémicos inducidos por una dieta enriquecida en colesterol. Los resultados in vitro permitieron comprobar que L. plantarum Y15 posee una elevada actividad BSH. In vivo, la suplementación con L. plantarum Y15 redujo el colesterol total y los triglicéridos en el suero y el hígado, aumentando el colesterol total y el ácido biliar total en las heces y reduciendo los cambios histopatológicos en el hígado. Además, la suplementación con L. plantarum Y15 moduló la microbiota intestinal al patrón del grupo de control, lo que implicó un aumento de la actividad de la BSH, que además influyó en la vía de señalización del FXR y el SHP, regulando al alza el nivel de expresión del CYP7A1 para mejorar el catabolismo del colesterol.
1. Introduction
Hypercholesterolemia is a leading risk factor for human cardiovascular disease (CVD), a major cause of death worldwide. The World Health Organization (WHO) delineated that high blood cholesterol is the cause of 2.6 million deaths and 29.7 million disability-adjusted life years annually worldwide (Mathers et al., Citation2009). Although statins are often well used to regulate cholesterol and reduced approximately 25% of CVD events, they are also questioned with undesirable side effects (Banach et al., Citation2015; Moss & Ramji, Citation2016). Therefore, it becomes crucial to find a healthy and effective way to lower cholesterol.
Probiotics are defined as “live microorganisms that impart a health benefit on the host, when consumed in sufficient amounts” (FAO/WHO, Citation2002). Increasing evidence demonstrates that probiotic bacteria have a cholesterol-lowering activity and can be considered a potential dietary therapy in preventing CVD. There are plenty of reports of the health benefits of L. plantarum, including cholesterol-lowering activity. It was found that L. plantarum CUL66 could reduce cholesterol in Caco-2 enterocytes (Michael et al., Citation2016). L. plantarum K21 has been proven to attenuate the plasma cholesterol effectively and triglyceride (TG) levels in diet-induced obese mice (DIO) mice (Wu et al., Citation2015). Cholesterol-lowering efficacy of L. plantarum CECT 7527, 7528, and 7529 in hypercholesterolaemic adults was reported (Fuentes et al., Citation2013). A clinical trial also showed that L. plantarum Q180 could ameliorate postprandial lipid metabolism and maintain a healthy intestinal environment (Y. E. Park et al., Citation2020).
There are multiple pathways to lower cholesterol by Lactobacillus strains. Firstly, Lactobacillus strains can assimilate cholesterol and catalyze bile salts’ hydrolysis by bile salt hydrolase (BSH) (Hosono, Citation1999; Jones et al., Citation2004). Secondly, Lactobacillus strains can converse cholesterol to coprostanol (Lye et al., Citation2010; Y.-H. Park et al., Citation2007). Thirdly, lipid metabolism is a complex process that involves the coordinated expression of numerous genes. Lactobacillus strains can regulate cholesterol homeostasis by modulating several gene expressions, such as transcription factors and genes related to cholesterol synthesis and decomposition (Kim et al., Citation2016). Fourthly, gut microbiota is associated with hypercholesterolemia (Prakash et al., Citation2011), and Lactobacillus strains can modulate gut microbiota. However, there is a lack of information about the cholesterol-lowering mechanisms of probiotics.
In this study, we selected a potential probiotic L. plantarum Y15 with high BSH activity and further investigated the effect and the mechanism of L. plantarum Y15 supplementation on hypercholesterolemia induced by a cholesterol-enriched diet. We also assessed the possible impact of this strain’s administration on the expression of several genes related to cholesterol metabolism in the liver. Three critical genes involved in bile acid biosynthesis are the Farnesoid X receptor (FXR), small heterodimer partner (SHP), and 7α-hydroxylase (CYP7A1). While the FXR is a bile salt deconjugation nuclear receptor, its activation triggers the SHP, a transcriptional suppressor that impairs the liver’s homolog-1 activity. These complex processes in decreased enzymatic CYP7A1 transcription that influences cholesterol metabolism in the liver (Kir et al., Citation2012; Parks et al., Citation1999; Pullinger et al., Citation2002). Furthermore, the cecal contents’ gut microbiota structure was analyzed to investigate the underlying correlation between the microbiota and cholesterol-lowering capacity. It is anticipated that this study will contribute to a more concrete theoretical understanding of the use of L. plantarum Y15 as a potential probiotic against hypercholesterolemia in functional foods and supplements formulation.
2. Materials and methods
2.1. Bacterial strain and culture
L. plantarum Y11, Y13, Y14, Y15 and Y17, L. fermentum Y21, Y22, Y24, Y26 and Y28, L. casei Y35, Y38 and Y39, L. acidophilus Y43, Y45 and Y47, L. sakei Y53, Y56 and Y58, and L. helveticus Y61 were isolated from yak yogurt in Gansu Province, China and identified by 16S rDNA similarity analysis. All strains with high acid and bile tolerances and adhesion ability were deposited at the School of Food and Biological Engineering, Zhengzhou University of Light Industry, China. All strains were anaerobically incubated in de Man, Rogosa, and Sharpe (MRS) broth (Hopebio Company, China) at 37°C for 24 h and sub-cultured twice before the experiment. Bacteria were harvested by centrifugation at 5000 g for 10 min. Cell pellets were then washed twice with sterile normal saline and resuspended at the desired concentration in normal saline for in vivo study.
2.2. Quantitative BSH activity
BSH activity was evaluated using ninhydrin assay (Dong et al., Citation2012). The substrate specificity was determined based on 5 mmol/L of glycodeoxycholic acid (GDCA, Sigma, USA) and taurodeoxycholic acid (TDCA, Sigma, USA), respectively. A standard curve of amino acid contents versus absorbance was prepared. The BSH activity (U/mg) was defined as the amount of enzyme that liberated 1 μmol of amino acids from the conjugated bile salts per minute. Specific activity was defined as units/mg of protein. Protein concentrations were determined following Lowry et al. (Lowry et al., Citation1951), and a standard curve was measured using bovine serum albumin (Sigma, USA).
2.3. Animals and experimental design
A total of 60 C57BL/6J mice (6 weeks old) were purchased from the Vital River Laboratory Animal Technology Company (Beijing, China) and housed in a room under controlled environmental conditions at 24 ± 2°C, relative humidity of 45–50% and with a 12-h light/dark cycle. The mice were placed in plastic cages for one week to acclimatize before the experiments and provided with a standard chow diet and water ad libitum. After acclimatization, mice were randomly divided into five groups. All groups were fed a cholesterol-enriched diet (consisted of 1% cholesterol, 0.3% bile salt, 10% lard, 5% yolk powder, and 83.7% normal chow diet) for four weeks to induce hypercholesterolemia (MC) except for the control group (NC), which received normal chow diet. In the meantime, the mice in the NC and MC groups were orally administered 0.2 mL saline solution once daily. The Y15 and Y61 groups were given 0.2 mL of saline containing 3 × 108 CFU of L. plantarum Y15 and L. helveticus Y61 by oral gavage once daily, respectively. The positive control group (PC) was orally treated with pravastatin (3 mg per kg body weight) once a day. All animal experiments reported in this study were approved by the Animal Care and Ethics Committee of Harbin Medical University. All diets were purchased from the Keo Synergy Feed Co. Ltd (TS1285, Beijing, China).
2.4. Determination of body weight and organ index
Mice weights were recorded every week. At the end of the dosing period, all animals were fasted for 16 h and then humanely sacrificed under diethyl ether anesthesia. The heart, liver, spleen, and kidney of mice were isolated and weighed for organ index calculation. The organ index was calculated using the following formula: organ index (%) = organ weight/body weight × 100.
2.5. Histopathological analysis of liver tissues
The liver was fixed in 10% neutral formalin for 48 h, after being dehydrated in graded ethanol, transparent in xylene, then embedded in paraffin wax. Tissue sections of 5 μm thickness were sliced and routinely stained with hematoxylin-eosin (HE). Histological differences between the groups were viewed and photographed with a light microscope.
2.6. Measurement of lipid profiles in serum, liver, and feces
Blood samples were drawn from the orbital venous plexus and centrifuged at 1150 g at 4°C for 10 min for the serum samples. Total fat in the liver tissues was extracted following Lepercq et al. (Citation2004). Feces were collected at the last two days, then dried and crushed into fecal powder, total cholesterol (TC) and total bile acid (TBA) were extracted from fecal powders previously reported (Fukada et al., Citation1991; Kajiura et al., Citation1998). The lipid profiles, including total cholesterol, triglyceride, low-density lipoprotein cholesterol (LDL-C), and high-density lipoprotein cholesterol (HDL-C) in serum, TC and TG in the liver, and TC and TBA in feces were measured by an automatic biochemistry analyzer (Toshiba, Tokyo, Japan) with enzyme assay kits.
2.7. Microbial analysis of cecal contents
Following standard instructions, we used the QIAamp DNA stool mini kit (Qiagen, Dusseldorf, Germany) to extract the total microbiota genomic DNA in the cecal contents from the NC, MC, and Y15 groups. The V4 region of the 16S rDNA were amplified by PCR. PCR amplicons were sequenced using the Illumina Miseq (Illumina, Santiago, USA). As earlier described by Edgar et al. (Citation2011), generated raw data were merged using the FLASH 1.2.7 software. We then discarded the chimera sequences using the UCHIME algorithm to obtain high-quality clean tags (Edgar et al., Citation2011). The Uparse software was used to convert clustered tags into distinct operational taxonomic units (OTUs) (Edgar, Citation2013). OTUs were classified by QIIME 1.7.0 (Caporaso et al., Citation2010). Linear Discriminant Analysis Effect Size (LEfSe) was used to performed with an effect size threshold of 2 (Stewart et al., Citation2017).
2.8. Real-time quantitative polymerase chain reaction (RT-qPCR) analyses of gene expression
Total RNA of each mouse’s liver tissue was extracted using Trizol reagent (Life Technologies, USA). According to the manufacturer’s instructions, cDNA was obtained by reverse transcription by using the PrimeScript RT reagent Kit with gDNA Eraser (Takara, Japan). The mRNA expression of genes was carried out by ABI 7500 fast real-time PCR system (Applied Biosystems, USA) using SYBR Premix Ex Taq (Takara, Japan). The primers (Comate Bioscience Co., Ltd, China) used in this study are presented in Table S1. Data analysis was performed by the 2−ΔΔCt method.
2.9. Statistical analysis
Data obtained from this research were from at least three independent assays, and the values reported were expressed as mean ± standard deviation (SD). Statistically, we used the one-way ANOVA method to compare datasets, followed by Duncan’s multiple range test, which were analyzed using SPSS 17.0 software (SPSS Inc., Chicago, IL, USA). This study measured statistical differences at the P < .05.
3. Results
3.1. BSH activity
The BSH activity of all tested strains to two different bile salts was determined and presented in . The strain L. helveticus Y61 did not deconjugate any of the conjugated bile acids. BSH activities had different substrate preferences and varied with the other strains, ranging from 0.123 to 1.803 U/mg and 0.027–0.517 U/mg for GDCA and TDCA hydrolase activities, respectively. L. plantarum Y15 possessed higher GDCA and TDCA hydrolase activities (1.803 ± 0.176 U/mg and 0.517 ± 0.055 U/mg) than the other tested strains. Based on the result of BSH activity, L. plantarum Y15 was the best candidate for alleviating hypercholesterolemia; thus it was selected for further in vivo experiments. L. helveticus Y61 was chosen as the negative strain.
Table 1. The BSH activity of Lactobacillus strains on bile salts.
Tabla 1. Actividad BSH de las cepas de Lactobacillus sobre las sales biliares
3.2. Effect of L. plantarum Y15 supplementation on body weight and organ index
As shown in , the mice’s body weight gain in the MC group was significantly higher than that of the NC group (P < .05). L. helveticus Y61 supplementation did not affect the bodyweight of cholesterol-enriched diet-fed mice. Conversely, it was effectively ameliorated after L. plantarum Y15 and pravastatin supplementations. No significant organ index changes, including kidney, heart, and spleen, were observed in any group. However, the cholesterol-enriched diet had a remarkable increase in the liver index than the NC group (P < .05), which implied a cholesterol-enriched diet could significantly affect hepatic tissues. L. plantarum Y15 and pravastatin supplementations could significantly alleviate liver index.
Table 2. Effect of Lactobacillus strains on body weight and organ index.
Tabla 2. Efecto de las cepas de Lactobacillus sobre el peso corporal y el índice de órganos
3.3. Effect of L. plantarum Y15 supplementation on liver histopathological changes
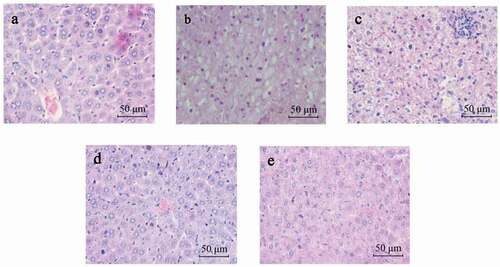
Normal liver histomorphology with a clear hepatocyte structure is presented in the NC group ()). On the other hand, liver specimens in the MC group were characterised by hepatic sinusoid stenosis, inflammation, swollen hepatocytes, diffuse fat vacuoles in the cytoplasm, and small focal necrotic nodules ()). Furthermore, results show that compared to the MC group, L. plantarum Y15 could effectively alleviate the liver histopathological variations, including relatively weak fat damage and unobserved small focal necrosis ()). L. helveticus Y61 was not as effective as that of L. plantarum Y15, which was similar to that observed in the PC group ()).
Figure 1. Representative photomicrographs of mice’s hepatic tissue (H&E staining; Scale bar, 50 μm). (a) NC group; (b) MC group; (c) Y61 group; (d) Y15 group; and (e) PC group.
Figura 1. Fotomicrografías representativas del tejido hepático de los ratones (tinción de H&E; barra de escala, 50 μm). (a) Grupo NC; (b) Grupo MC; (c) Grupo Y61; (d) Grupo Y15; y (e) Grupo PC
3.4. Effect of L. plantarum Y15 supplementation on lipid profiles in hypercholesterolemic mice
The lipid profiles in serum, liver, and feces are shown in . Increased serum lipid profiles confirmed hypercholesterolemia after four weeks of cholesterol-enriched diet feeding. As shown in ), TC, TG, and LDL-C levels in the MC group were significantly elevated than in the NC group (P < .05). However, there was no remarkable change in HDL-C level in all groups. There was no difference in TC, TG and LDL-C levels between the Y61 and MC groups (P > .05), suggesting that L. helveticus Y61 supplementation did not mitigate serum lipid levels in mice with hypercholesteremia. However, these parameters in the Y15 and PC groups were effectively ameliorated, indicating that L. plantarum Y15 supplementation could effectively reduce hypercholesteremia.
Figure 2. Comparison of lipid profiles in serum, liver, and feces among experimental groups of mice. (a) Lipid profiles in serum; (b) Lipid profiles in the liver; and (c) Lipid profiles in feces. TC, total cholesterol; TG, triglyceride; LDL-C, low-density lipoprotein cholesterol; HDL-C, high-density lipoprotein cholesterol; and TBA, total bile acid. Values are mean ± SD (n = 6 independent experiment). Significant differences (P < .05) are indicated with different letters above the graphical bars.
Figura 2. Comparación de perfiles lipídicos en suero, hígado y heces entre los grupos experimentales de ratones. (a) Perfiles lipídicos en suero; (b) Perfiles lipídicos en el hígado; y (c) Perfiles lipídicos en las heces. CT, colesterol total; TG, triglicéridos; LDL-C, colesterol de lipoproteínas de baja densidad; HDL-C, colesterol de lipoproteínas de alta densidad; y TBA, ácido biliar total. Los valores son la media ± DE (n = 6 experimentos independientes). Las diferencias significativas (P < .05) se indican con letras diferentes sobre las barras gráficas
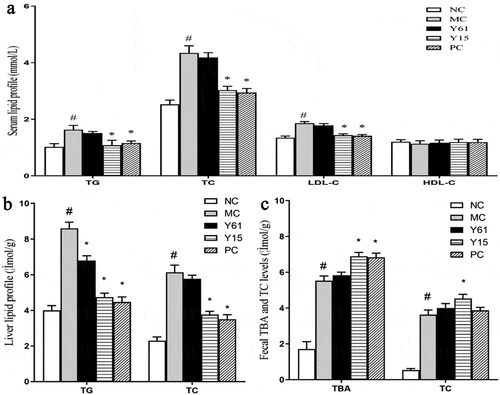
As shown in ), cholesterol-enriched diet feeding elevated TG and TC levels in the liver compared with the NC group. L. helveticus Y61 supplementation only significantly lowered the TG level, whereas significant decreases in TC and TG levels were found in the Y15 group, which showed the same trend with the PC group.
The TBA and TC levels in feces in the MC group were higher than those in the NC group, the Y15 group showed an increased effect on these two parameters ()), indicating that L. plantarum Y15 supplementation could lower cholesterol level via the excretion of TC and TBA. However, no difference in TBA and TC levels was observed between the Y61 and MC groups.
As aforementioned, L. plantarum Y15 supplementation had a significant cholesterol-lowering effect. Therefore, the Y15 group was selected for further study to investigate possible mechanisms of action.
3.5. Effect of L. plantarum Y15 supplementation on gut microbiota
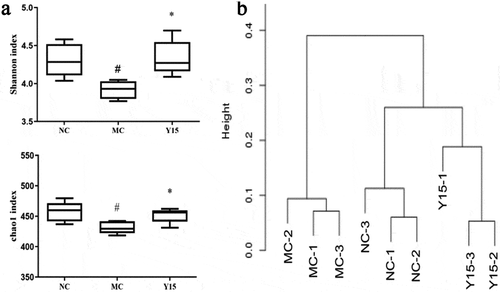
This study examined the gut microbiota of the NC, MC, and Y15 groups using 16S rRNA gene amplicon sequencing. To illustrate whether gut microbiota alteration was related with the prevent effects of L. plantarum Y15 supplementation on hypercholesterolemia. Among the bacterial α-diversity measurements, the Shannon index and Chao 1 index were used to estimate gut microbiota’s richness and diversity. As shown in ), the Shannon index and Chao 1 index in the Y15 group were remarkably higher than those in the MC group (P < .05), while no apparent differences with the NC group were observed (P > .05). This finding implied that L. plantarum Y15 supplementation could elevate the diversity of the gut microbiota. Also, the hierarchical clustering tree of Bray-Curtis distances showed that the Y15 group have a better correlation with the NC group than that with the MC group ()). This finding indicated that L. plantarum Y15 supplementation modulated the gut microbiota’s structure in the MC group to that of the NC group.
Figure 3. The effect of L. plantarum Y15 supplementation on gut microbiota α-diversity and β-diversity. (a) Shannon and Chao 1 index; and (b) Hierarchical clustering tree of Bray-Curtis distances. Values are mean ± SD (n = 3 independent experiment). # P < .05: significantly different compared with the NC group; * P < .05: significantly different compared with the MC group.
Figura 3. Efecto de la suplementación con L. plantarum Y15 en la diversidad α y β de la microbiota intestinal. (a) Índice de Shannon y Chao 1; y (b) Árbol de agrupación jerárquica de las distancias Bray-Curtis. Los valores son la media ± DE (n = 3 experimentos independientes). # P < .05: significativamente diferente en comparación con el grupo NC; * P < .05: significativamente diferente en comparación con el grupo MC
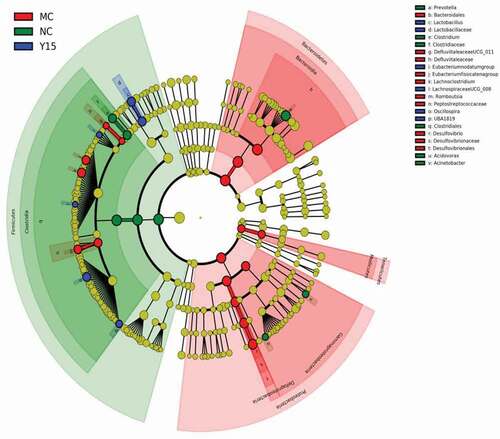
As shown in ), the predominant microbiota in all groups was Bacteroidetes, Firmicutes, and Proteobacteria at the phylum level. After four weeks of cholesterol-enriched diet feeding, the abundance of the phyla Firmicutes was higher in the NC (41.80%) and Y15 (47.76%) groups than that in the MC group (29.85%). In contrast, the abundance of Bacteroidetes was lower in the NC (41.22%) and Y15 groups (34.62%) than that in the MC group (51.45%). Proteobacteria’s relative abundance increased from 11.16% to 15.73% after the cholesterol-enriched diet feeding, and L. plantarum Y15 supplementation alleviated the increase in Proteobacteria (13.54%).
Figure 4. L. plantarum Y15 supplementation manipulated the gut microbiota composition in hypercholesterolemic mice induced by cholesterol-enriched diet fed. (a) Gut microbiota distribution at the phylum level; and (b) Gut microbiota distribution at the genus level. Values are mean ± SD (n = 3 independent experiment). # P < .05: significantly different compared with the NC group; * P < .05: significantly different compared with the MC group.
Figura 4. La suplementación con L. plantarum Y15 manipuló la composición de la microbiota intestinal en ratones hipercolesterolémicos inducidos por una dieta enriquecida en colesterol. (a) Distribución de la microbiota intestinal a nivel de filo; y (b) Distribución de la microbiota intestinal a nivel de género. Los valores son la media ± DE (n = 3 experimentos independientes). # P < .05: significativamente diferente en comparación con el grupo NC; * P < .05: significativamente diferente en comparación con el grupo MC
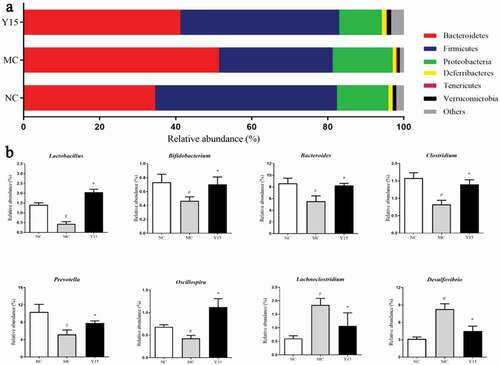
From the insight of genus level, the genera abundance varied in the NC, MC and Y15 groups ()). The relative abundances of Lactobacillus, Bifidobacterium, Bacteroides, Clostridium, Prevotella, and Oscillospira were decreased in the MC group, when compared to the NC group, all of which substantially improved following L. plantarum Y15 supplementation. Furthermore, some elevated genera in the MC group, such as Lachnoclostridium and Desulfovibrio, were inhibited by L. plantarum Y15 supplementation.
LEfSe analysis was used to identify the key phylotypes responsible for the differences biomarkers between groups. As shown in , Desulfovibrio and Lachnoclostridium were significantly more abundant in the MC group, Clostridium and Prevotella were enriched in the NC group, Lactobacillus and Oscillospira were significantly more abundant in the Y15 group. LEfSe analyses were consistent with the aforementioned results.
3.6. Effect of L. plantarum Y15 supplementation on gene expression related to hepatic cholesterol homeostasis
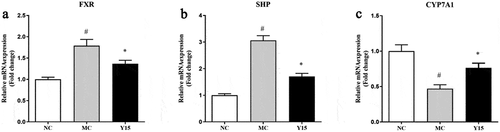
To further investigate the cholesterol-lowering effect mechanism, the expression levels of genes related to cholesterol metabolism were analyzed by qRT-PCR. As shown in , cholesterol-enriched diet feeding for four weeks upregulated FXR and SHP genes’ expression levels. However, it downregulated the expression of CYP7A1. The expression levels of FXR and SHP genes were inhibited by L. plantarum Y15 supplementation. The expression level of the CYP7A1 gene was significantly upregulated in the Y15 group relative to the MC group. These findings indicated that L. plantarum Y15 supplementation could lower cholesterol by increasing cholesterol decomposition genes’ expression levels.
Figure 6. Effect of L. plantarum Y15 supplementation on the expression levels of genes related to cholesterol metabolism in the liver. (A) FXR; (b) SHP; and (c) CYP7A1. Values are mean ± SD (n = 6 independent experiment). # P < .05: significantly different compared with the NC group; * P < .05: significantly different compared with the MC group.
Figura 6. Efecto de la suplementación con L. plantarum Y15 sobre los niveles de expresión de los genes relacionados con el metabolismo del colesterol en el hígado. (a) FXR; (b) SHP; y (c) CYP7A1. Los valores son la media ± DE (n = 6 experimentos independientes). # P < .05: significativamente diferente en comparación con el grupo NC; * P < .05: significativamente diferente en comparación con el grupo MC
4. Discussion
Hypercholesterolemia is a leading risk factor for the development of CVD. It has been advocated that probiotics may be a potential supplemental tool for hypercholesterolemia (Anandharaj et al., Citation2014). As reported, BSH-deficient mutant-Lactobacillus strain did not significantly impact the hepatic cholesterol-lowering action in hamsters (C. F. Guo et al., Citation2018). Lactobacillus strain overexpression of BSH has beneficial effects against hypercholesterolemia (Wang et al., Citation2019). BSH activity in vitro was a crucial index for screening probiotics with potential ability against hypercholesterolemia in vivo (Tsai et al., Citation2014; Zhang et al., Citation2017). Thus, L. plantarum Y15 with higher BSH activity in vitro was selected from twenty Lactobacillus strains as a potential candidate for alleviating hypercholesterolemia.
Furthermore, this strain was further studied in vivo in hypercholesterolemic mice induced by cholesterol-enriched diet feeding for four weeks. The intake of a cholesterol-enriched diet increased the liver index and elevated the TC and TG contents in mice’s liver, consistent with the diffuse fat vacuoles in liver histopathological sections observed in the MC group. However, L. plantarum Y15 supplementation could significantly reduce the liver index, the TC and TG contents and improve liver histopathological changes. These results agreed with the earlier reports in which BSH-active L. plantarum KLDS 1.0344 could improve hepatic cholesterol metabolism in rats fed a high-cholesterol diet (L. Guo et al., Citation2019). These findings implied that L. plantarum Y15 had the cholesterol-lowering ability in vivo by regulating cholesterol metabolism.
Accumulating evidence suggests an interactive link between gut microbiota and hypercholesterolemia (Caparrós-Martín et al., Citation2017; Khan et al., Citation2018). L. plantarum Y15 supplementation decreased Lachnoclostridium and Desulfovibrio levels. Supporting this, similar variations were also found in the hypercholesterolemia mice after L. plantarum H6 treatment (Qu et al., Citation2020). Lachnoclostridium belonged to the Lachnospiraceae family, has been shown to have a positive correlation with body weight gain in DIO mice (Ravussin et al., Citation2012). Desulfovibrio is one factor that mediates the genetic mechanism of diabetes (Zhu et al., Citation2015). In contrast, upregulation of the relative abundances of Prevotella, Oscillospira, Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium was found in the Y15 group. Some previous studies indicated that the abundance of Prevotella was decreased in patients with non-alcoholic fatty liver disease, and Oscillospira is negatively correlated with serum cholesterol concentration (Lye et al., Citation2017; Targher & Byrne, Citation2017). Lactobacillus and Bifidobacterium are well-known probiotics with several beneficial effects on body metabolism (Li et al., Citation2018). The increased proportion of Lactobacillus may be attributed to L. plantarum Y15 survival in the intestine with high resistance to the simulated gastrointestinal environment (Zhang, Citation2006). It has been reported that Bacteroides fragilis-intestinal FXR axis could improve hyperglycemia. Clostridium can produce butyrate, which plays a vital role in lowering cholesterol (Ishimwe et al., Citation2015; Rivera-Chávez et al., Citation2016). However, its abundance was reduced in the atorvastatin-treated group (Khan et al., Citation2018). Above all, the presence of BSH was demonstrated in Lactobacillus, Bifidobacterium, Bacteroides and Clostridium (Horáčková et al., Citation2018).
BSH can hydrolyse conjugated bile salts to liberate deconjugated bile acids, showing the protective effect in lowering serum cholesterol levels by dual pathways. Firstly, deconjugated bile acids are less soluble and less efficiently reabsorbed in the gut tract than conjugated bile salts (Begley et al., Citation2006; Nguyen et al., Citation2007). Thus, they are excreted in feces and subsequently synthesized de novo from hepatic cholesterol, thereby decreasing their serum levels (Lye et al., Citation2010). In agreement with previous reports by Ding et al. (Citation2017), the elevated level of TBA along with TC in feces and the decreased level of TC accompanied with TG and LDL-C in serum were found after L. plantarum Y15 supplementation.
Secondly, deconjugated bile acids can activate the nuclear receptor, FXR, which regulates the homeostasis of bile acid, lipid, and glucose metabolism (Degirolamo et al., Citation2014). Therefore, the FXR activity can be influenced depending on the bile acid pool profile modified by BSH. It has been demonstrated that decreased hepatic FXR-SHP signaling increased catabolism of cholesterol and biosynthesis of bile acids by CYP7A1 (de Aguiar Vallim et al., Citation2013; Jones et al., Citation2013), which is the rate-limiting enzyme in bile acid synthesis in the liver (Pullinger et al., Citation2002). Our study confirmed these findings in that the mRNA expression levels of FXR and SHP were decreased by L. plantarum Y15 supplementation. In contrast, the hepatic expression level of CYP7A1 in the Y15 group was significantly down-regulated compared with the MC group. These results were in line with the functions of L. plantarum KCTC3928 (Jeun et al., Citation2010) and L. casei WQH01 (overexpression of bsh1) (Wang et al., Citation2019).
In summary, L. plantarum Y15 could increase the relative abundances of some genera of gut bacteria, such as Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium, with BSH activity. The hydrolyzation released deconjugated bile acids with low resorption, leading to a reduction in the flow to the liver through enterohepatic circulation. Subsequently, it significantly decreased the liver’s FXR expression level, which downregulated the expression level of SHP and upregulated the expression level of CYP7A1. Finally, the upregulated CYP7A1 elevated the cholesterol catabolism. These findings indicated that L. plantarum Y15 plays a crucial role in regulating cholesterol and bile acids homeostasis via the gut microbiota- bile acids interaction in mice fed cholesterol-enriched diets.
5. Conclusion
L. plantarum Y15 demonstrated a high BSH activity in vitro. L. plantarum Y15 showed the protective effect against hypercholesterolemia in vivo by improving the serum, liver, and feces’ lipid profiles and relieving hepatic pathological damages. Moreover, L. plantarum Y15 supplementation regulated the gut microbiota with BSH activity, resulting in upregulation of deconjugated bile acids with low resorption, which could further influence the hepatic FXR-SHP signaling pathway to upregulate the expression level of CYP7A1 to enhance the cholesterol catabolism.
Supplemental Material
Download MS Word (22.1 KB)Disclosure statement
No potential conflict of interest was reported by the author(s).
Supplementary material
Supplemental data for this article can be accessed on the publisher’s website
Additional information
Funding
References
- Anandharaj, M., Sivasankari, B., & Parveen Rani, R. (2014). Effects of probiotics, prebiotics, and synbiotics on hypercholesterolemia: A review. Chinese Journal of Biology,2014,(2014-2-27), 2014(3–4), 1–7. https://doi.org/https://doi.org/10.1155/2014/572754
- Banach, M., Rizzo, M., Toth, P. P., Farnier, M., Davidson, M. H., Alrasadi, K., Aronow, W. S., Athyros, V., Djuric, D. M., Ezhov, M. V., Greenfield, R. S., Hovingh, G. K., Kostner, K., Serban, C., Lighezan, D., Fras, Z., Moriarty, P. M., Muntner, P., Goudev, A., Ceska, R., & Mikhailidis, D. P. (2015). Statin intolerance – An attempt at a unified definition. Position paper from an International Lipid Expert Panel. Archives of Medical Science Ams, 11(1), 1. https://doi.org/https://doi.org/10.5114/aoms.2015.49807
- Begley, M., Hill, C., & Gahan, C. G. (2006). Bile salt hydrolase activity in probiotics. Applied and Environmental Microbiology, 72(3), 1729–1738. https://doi.org/https://doi.org/10.1128/AEM.72.3.1729-1738.2006
- Caparrós-Martín, J. A., Lareu, R. R., Ramsay, J. P., Peplies, J., Reen, F. J., Headlam, H. A., Ward, N. C., Croft, K. D., Newsholme, P., & Hughes, J. D. (2017). Statin therapy causes gut dysbiosis in mice through a PXR-dependent mechanism. Microbiome, 5(1), 95. https://doi.org/https://doi.org/10.1186/s40168-017-0312-4
- Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., Fierer, N., Peña, A. G., Goodrich, J. K., Gordon, J. I., Huttley, G. A., Kelley, S. T., Knights, D., Koenig, J. E., Ley, R. E., Lozupone, C. A., McDonald, D., Muegge, B. D., Pirrung, M., Reeder, J., & Knight, R. (2010). QIIME allows analysis of high-throughput community sequencing data. Nature Methods, 7(5), 335–336. https://doi.org/https://doi.org/10.1038/nmeth.f.303
- de Aguiar Vallim, T. Q., Tarling, E. J., & Edwards, P. A. (2013). Pleiotropic roles of bile acids in metabolism. Cell Metabolism, 17(5), 657–669. https://doi.org/https://doi.org/10.1016/j.cmet.2013.03.013
- Degirolamo, C., Rainaldi, S., Bovenga, F., Murzilli, S., & Moschetta, A. (2014). Microbiota modification with probiotics induces hepatic bile acid synthesis via downregulation of the Fxr-Fgf15 axis in mice. Cell Reports, 7(1), 12–18. https://doi.org/https://doi.org/10.1016/j.celrep.2014.02.032
- Ding, W., Shi, C., Chen, M., Zhou, J., Long, R., & Guo, X. (2017). Screening for lactic acid bacteria in traditional fermented Tibetan yak milk and evaluating their probiotic and cholesterol-lowering potentials in rats fed a high-cholesterol diet. Journal of Functional Foods, 32(2017), 324–332. https://doi.org/https://doi.org/10.1016/j.jff.2017.03.021
- Dong, Z., Zhang, J., Lee, B., Li, H., Du, G., & Chen, J. (2012). A bile salt hydrolase gene of Lactobacillus plantarum BBE7 with high cholesterol-removing activity. European Food Research and Technology, 235(3), 419–427. https://doi.org/https://doi.org/10.1007/s00217-012-1769-9
- Edgar, R. C. (2013). UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nature Methods, 10(10), 996. https://doi.org/https://doi.org/10.1038/nmeth.2604
- Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., & Knight, R. (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics, 27(16), 2194–2200. https://doi.org/https://doi.org/10.1093/bioinformatics/btr381
- FAO/WHO. (2002). Joint FAO/WHO working group report on drafting guidelines for the evaluation of probiotics in food.
- Fuentes, M. C., Lajo, T., Carrión, J. M., & Cuñé, J. (2013). Cholesterol-lowering efficacy of Lactobacillus plantarum CECT 7527, 7528 and 7529 in hypercholesterolaemic adults. British Journal of Nutrition, 109(10), 1866–1872. https://doi.org/https://doi.org/10.1017/S000711451200373X
- Fukada, Y., Kimura, K., & Ayaki, Y. (1991). Effect of chitosan feeding on intestinal bile acid metabolism in rats. Lipids, 26(5), 395–399. https://doi.org/https://doi.org/10.1007/BF02537206
- Guo, C. F., Zhang, S., Yuan, Y. H., Li, J. Y., & Yue, T. L. (2018). Bile salt hydrolase and S‐layer protein are the key factors affecting the hypocholesterolemic activity of Lactobacillus casei-fermented milk in hamsters. Molecular Nutrition & Food Research, 62(24), 1800728. https://doi.org/https://doi.org/10.1002/mnfr.201800728
- Guo, L., Wang, L., Liu, F., Li, B., Tang, Y., Yu, S., Zhang, D., & Huo, G. (2019). Effect of bile salt hydrolase-active Lactobacillus plantarum KLDS 1.0344 on cholesterol metabolism in rats fed a high-cholesterol diet. Journal of Functional Foods, 61(2019), 103497. https://doi.org/https://doi.org/10.1016/j.jff.2019.103497
- Horáčková, Š., Plocková, M., & Demnerová, K. (2018). Importance of microbial defence systems to bile salts and mechanisms of serum cholesterol reduction. Biotechnology Advances, 36(3), 682–690. https://doi.org/https://doi.org/10.1016/j.biotechadv.2017.12.005
- Hosono, A. (1999). Bile tolerance, taurocholate deconjugation, and binding of cholesterol by Lactobacillus gasseri strains. Journal of Dairy Science, 82(2), 243–248. https://doi.org/https://doi.org/10.3168/jds.S0022-0302(99)75229-X
- Ishimwe, N., Daliri, E. B., Lee, B. H., Fang, F., & Du, G. (2015). The perspective on cholesterol‐lowering mechanisms of probiotics. Molecular Nutrition & Food Research, 59(1), 94–105. https://doi.org/https://doi.org/10.1002/mnfr.201400548
- Jeun, J., Kim, S., Cho, S.-Y., Jun, H.-J., Park, H.-J., Seo, J.-G., Chung, M.-J., & Lee, S.-J. (2010). Hypocholesterolemic effects of Lactobacillus plantarum KCTC3928 by increased bile acid excretion in C57BL/6 mice. Nutrition, 26(3), 321–330. https://doi.org/https://doi.org/10.1016/j.nut.2009.04.011
- Jones, M. L., Chen, H., Ouyang, W., Metz, T., & Prakash, S. (2004). Microencapsulated genetically engineered Lactobacillus plantarum 80 (pCBH1) for bile acid deconjugation and its implication in lowering cholesterol. BioMed Research International, 2004(1), 61–69. https://doi.org/https://doi.org/10.1155/S1110724304307011
- Jones, M. L., Tomaro-Duchesneau, C., Martoni, C. J., & Prakash, S. (2013). Cholesterol lowering with bile salt hydrolase-active probiotic bacteria, mechanism of action, clinical evidence, and future direction for heart health applications. Expert Opinion on Biological Therapy, 13(5), 631–642. https://doi.org/https://doi.org/10.1517/14712598.2013.758706
- Kajiura, K., Ohkusa, T., & Okayasu, I. (1998). Relationship between fecal bile acids and the occurrence of colorectal neoplasia in experimental murine ulcerative colitis. Digestion, 59(1), 69–72. https://doi.org/https://doi.org/10.1159/000007469
- Khan, T. J., Ahmed, Y. M., Zamzami, M. A., Mohamed, S. A., Khan, I., Baothman, O. A., Mehanna, M. G., & Yasir, M. (2018). Effect of atorvastatin on the gut microbiota of high fat diet-induced hypercholesterolemic rats. Scientific Reports, 8(1), 662. https://doi.org/https://doi.org/10.1038/s41598-017-19013-2
- Kim, B., Park, K. Y., Ji, Y., Park, S., Holzapfel, W., & Hyun, C. K. (2016). Protective effects of Lactobacillus rhamnosus GG against dyslipidemia in high-fat diet-induced obese mice. Biochemical & Biophysical Research Communications, 473(2), 530–536. https://doi.org/https://doi.org/10.1016/j.bbrc.2016.03.107
- Kir, S., Zhang, Y., Gerard, R. D., Kliewer, S. A., & Mangelsdorf, D. J. (2012). Nuclear receptors HNF4α and LRH-1 cooperate in regulating Cyp7a1 in vivo. Journal of Biological Chemistry, 287(49), 41334–41341. https://doi.org/https://doi.org/10.1074/jbc.M112.421834
- Lepercq, P., Relano, P., Cayuela, C., & Juste, C. (2004). Bifidobacterium animalis strain DN-173 010 hydrolyses bile salts in the gastrointestinal tract of pigs. Scandinavian Journal of Gastroenterology, 39(12), 1266–1271. https://doi.org/https://doi.org/10.1080/00365520410003515
- Li, B., Evivie, S. E., Lu, J., Jiao, Y., Wang, C., Li, Z., Liu, F., & Huo, G. (2018). Lactobacillus helveticus KLDS1. 8701 alleviates D-galactose-induced aging by regulating Nrf-2 and gut microbiota in mice. Food & Function, 9(12), 6586–6598. https://doi.org/https://doi.org/10.1039/C8FO01768A
- Lowry, O. H., Rosebrough, N. J., Farr, A. L., & Randall, R. J. (1951). Protein measurement with the Folin phenol reagent. Journal of Biological Chemistry, 193(1), 265–275. https://doi.org/https://doi.org/10.1016/S0021-9258(19)52451-6
- Lye, H.-S., Kato, T., Low, W.-Y., Taylor, T. D., Prakash, T., Lew, L.-C., Ohno, H., & Liong, M.-T. (2017). Lactobacillus fermentum FTDC 8312 combats hypercholesterolemia via alteration of gut microbiota. Journal of Biotechnology, 262(2017), 75–83. https://doi.org/https://doi.org/10.1016/j.jbiotec.2017.09.007
- Lye, H.-S., Rahmat-Ali, G. R., & Liong, M.-T. (2010). Mechanisms of cholesterol removal by lactobacilli under conditions that mimic the human gastrointestinal tract. International Dairy Journal, 20(3), 169–175. https://doi.org/https://doi.org/10.1016/j.idairyj.2009.10.003
- Lye, H.-S., Rusul, G., & Liong, M.-T. (2010). Removal of cholesterol by lactobacilli via incorporation and conversion to coprostanol. Journal of Dairy Science, 93(4), 1383–1392. https://doi.org/https://doi.org/10.3168/jds.2009-2574
- Mathers, C., Stevens, G., & Mascarenhas, M. (2009). Global health risks: Mortality and burden of disease attributable to selected major risks. World Health Organization.
- Michael, D., Moss, J., Calvente, D. L., Garaiova, I., Plummer, S., & Ramji, D. (2016). Lactobacillus plantarum CUL66 can impact cholesterol homeostasis in Caco-2 enterocytes. Beneficial Microbes, 7(3), 443–451. https://doi.org/https://doi.org/10.3920/BM2015.0146
- Moss, J. W., & Ramji, D. P. (2016). Nutraceutical therapies for atherosclerosis. Nature Reviews Cardiology, 13(9), 513. https://doi.org/https://doi.org/10.1038/nrcardio.2016.103
- Nguyen, T., Kang, J., & Lee, M. (2007). Characterization of Lactobacillus plantarum PH04, a potential probiotic bacterium with cholesterol-lowering effects. International Journal of Food Microbiology, 113(3), 358–361. https://doi.org/https://doi.org/10.1016/j.ijfoodmicro.2006.08.015
- Park, Y. E., Kim, M. S., Shim, K. W., Kim, Y.-I., Chu, J., Kim, B.-K., Choi, I. S., & Kim, J. Y. (2020). Effects of Lactobacillus plantarum Q180 on postprandial lipid levels and intestinal environment: A double-blind, randomized, placebo-controlled, parallel trial. Nutrients, 12(1), 255. https://doi.org/https://doi.org/10.3390/nu12010255
- Park, Y.-H., Kim, J.-G., Shin, Y.-W., Kim, S.-H., & Whang, K.-Y. (2007). Effect of dietary inclusion of Lactobacillus acidophilus ATCC 43121 on cholesterol metabolism in rats. Journal of Microbiology and Biotechnology, 17(4), 655–662. https://doi.org/https://doi.org/10.1007/s10295-006-0202-4
- Parks, D. J., Blanchard, S. G., Bledsoe, R. K., Chandra, G., Consler, T. G., Kliewer, S. A., Stimmel, J. B., Willson, T. M., Zavacki, A. M., & Moore, D. D. (1999). Bile acids: Natural ligands for an orphan nuclear receptor. Science, 284(5418), 1365–1368. https://doi.org/https://doi.org/10.1126/science.284.5418.1365
- Prakash, S., Tomaro-Duchesneau, C., Saha, S., & Cantor, A. (2011). The gut microbiota and human health with an emphasis on the use of microencapsulated bacterial cells. BioMed Research International, 2011, Article ID 981214. https://doi.org/https://doi.org/10.1155/2011/981214
- Pullinger, C. R., Eng, C., Salen, G., Shefer, S., Batta, A. K., Erickson, S. K., Verhagen, A., Rivera, C. R., Mulvihill, S. J., & Malloy, M. J. (2002). Human cholesterol 7α-hydroxylase (CYP7A1) deficiency has a hypercholesterolemic phenotype. The Journal of Clinical Investigation, 110(1), 109–117. https://doi.org/https://doi.org/10.1172/JCI0215387
- Qu, T., Yang, L., Wang, Y., Jiang, B., Shen, M., & Ren, D. (2020). Reduction of serum cholesterol and its mechanism by Lactobacillus plantarum H6 screened from local fermented food products. Food & Function, 11(2), 1397–1409. https://doi.org/https://doi.org/10.1039/C9FO02478F
- Ravussin, Y., Koren, O., Spor, A., Leduc, C., Gutman, R., Stombaugh, J., Knight, R., Ley, R. E., & Leibel, R. L. (2012). Responses of gut microbiota to diet composition and weight loss in lean and obese mice. Obesity, 20(4), 738–747. https://doi.org/https://doi.org/10.1038/oby.2011.111
- Rivera-Chávez, F., Zhang, L. F., Faber, F., Lopez, C. A., Byndloss, M. X., Olsan, E. E., Xu, G., Velazquez, E. M., Lebrilla, C. B., & Winter, S. E. (2016). Depletion of butyrate-producing Clostridia from the gut microbiota drives an aerobic luminal expansion of Salmonella. Cell Host & Microbe, 19(4), 443–454. https://doi.org/https://doi.org/10.1016/j.chom.2016.03.004
- Stewart, C. J., Embleton, N. D., Marrs, E. C. L., Smith, D. P., Fofanova, T., Nelson, A., Skeath, T., Perry, J. D., Petrosino, J. F., & Berrington, J. E. (2017). Longitudinal development of the gut microbiome and metabolome in preterm neonates with late onset sepsis and healthy controls. Microbiome, 5(1), 75. https://doi.org/https://doi.org/10.1186/s40168-017-0295-1
- Targher, G., & Byrne, C. D. (2017). Non-alcoholic fatty liver disease: An emerging driving force in chronic kidney disease. Nature Reviews Nephrology, 13(5), 297. https://doi.org/https://doi.org/10.1038/nrneph.2017.16
- Tsai, -C.-C., Lin, -P.-P., Hsieh, Y.-M., Zhang, Z.-Y., Wu, H.-C., & Huang, -C.-C. (2014). Cholesterol-lowering potentials of lactic acid bacteria based on bile-salt hydrolase activity and effect of potent strains on cholesterol metabolism in vitro and in vivo. The Scientific World Journal, 2014(3), 1–10. https://doi.org/https://doi.org/10.1155/2014/690752
- Wang, G., Huang, W., Xia, Y., Xiong, Z., & Ai, L. (2019). Cholesterol-lowering potentials of Lactobacillus strain overexpression of bile salt hydrolase on high cholesterol diet-induced hypercholesterolemic mice. Food & Function, 10(3), 1684–1695. https://doi.org/https://doi.org/10.1039/C8FO02181C
- Wu, -C.-C., Weng, W.-L., Lai, W.-L., Tsai, H.-P., Liu, W.-H., Lee, M.-H., & Tsai, Y.-C. (2015). Effect of Lactobacillus plantarum strain K21 on high-fat diet-fed obese mice. Evidence-Based Complementary and Alternative Medicine, 2015(1), 1–9. https://doi.org/https://doi.org/10.1155/2015/391767
- Zhang, D. D. (2006). Mechanistic studies of the Nrf2-Keap1 signaling pathway. Drug Metabolism Reviews, 38(4), 769–789. https://doi.org/https://doi.org/10.1080/03602530600971974
- Zhang, F., Qiu, L., Xu, X., Liu, Z., Zhan, H., Tao, X., Shah, N. P., & Wei, H. (2017). Beneficial effects of probiotic cholesterol-lowering strain of Enterococcus faecium WEFA23 from infants on diet-induced metabolic syndrome in rats. Journal of Dairy Science, 100(3), 1618–1628. https://doi.org/https://doi.org/10.3168/jds.2016-11870
- Zhu, M., Kang, Y., & Du, M. (2015). Maternal obesity alters gut microbial ecology in offspring of NOD mice. The FASEB Journal, 29(1_Suppl), 105.103. https://doi.org/https://doi.org/10.1155/2015/391767